Drawing 2026-05-01 12.16.40.excalidraw
⚠ Switch to EXCALIDRAW VIEW in the MORE OPTIONS menu of this document. ⚠ You can decompress Drawing data with the command palette: ‘Decompress current Excalidraw file’. For more info check in plugin settings under ‘Saving’
Excalidraw Data
Text Elements
Link to original
TARGET DECK: MED::I::Signaling Pathways in Health and Disease::Metabolic Biochemistry::02 - Oxidative Phosphorylation
- integrati appunti marco
Bioenergetics
Mechanisms of ATP Synthesis — Phosphorylation of ADP
Substrate-Level Phosphorylation
Mechanism:
Two types of phosphorylation
- Substrate-level: direct transfer of a phosphate group from a high-energy intermediate to ADP
- Oxidative: coupled to the electron transfer chain
Example: GA3P → 1,3BPG
The oxidation of GA3P is very exergonic: the energy is used to incorporate to form the high-energy bond of 1,3BPG.
Two steps:
- Oxidation of the aldehyde to a carboxylic group
- Attachment of
Primary Function of Mitochondria
What is the primary function of mitochondria?
Mitochondria transfer electrons to O₂ via the respiratory chain, producing H₂O and using that energy to synthesize ATP by oxidative phosphorylation.
Core function
Mitochondria transfer electrons (from oxidative metabolism intermediates) to , producing via the Respiratory Chain, and harness that energy to synthesize ATP via Oxidative Phosphorylation.
Additional Functions of Mitochondria
- Most catabolic pathways (TCA cycle, fatty acid -oxidation, amino acid oxidation)
- Some biosynthetic pathways or reactions
- Mitochondrial DNA and protein synthesis
- Generation of Reactive Oxygen Species (ROS)
- Involved in signal transduction networks
- Involved in apoptosis (programmed cell death)
Respiratory Chain
A sequence of redox coenzymes that carries electrons from substrates to oxygen.
Overall reaction:
The respiratory chain:
Why multiple steps?
By subdividing the energy loss into several steps, energy can be conserved in the form of ATP chemical bonds — rather rather than lost as heat in a single reaction.
What's ?
SH₂is a generic notation for a reduced substrate (an electron/hydrogen donor).
S = any oxidizable substrate
H₂ = the two hydrogen atoms (electrons + protons) being removed from it
For example:
- NADH → here SH₂ would be the metabolite that donated H₂ to NAD⁺ to form NADH (e.g., malate, isocitrate from the TCA cycle)
So the reaction
SH₂ → Ssimply means the substrate gets oxidized by losing two hydrogens, which then enter the respiratory chain.
Summary
- Structure of Mitochondria
- The Respiratory Chain
- ATP Synthase
- Chemio-Osmotic Coupling
- Respiratory Control and Uncoupling
- Mitochondrial DNA
- Mitochondria and Reactive Oxygen Species (ROS)
Mitochondrial Structure
- Outer membrane: permeable to small molecules and ions.
- Inner membrane: impermeable to small molecules and ions (requires specific transporters).
- Mitochondrial matrix: contains enzymes of carbohydrate (downstream PDH), fatty acids, and amino acid catabolism.
Inner membrane impermeability
The impermeability of the inner mitochondrial membrane is essential — it allows the proton gradient to build up and drive ATP synthesis.
Anki cloze
The inner mitochondrial membrane is impermeable to {1:small molecules and ions}, which allows the {1:proton gradient} to build up and drive ATP synthesis.
Mitochondrial Respiratory Chain
Coenzymes of the Respiratory Chain
5 types of electron carriers
- ; (flavoproteins)
- (ubiquinone / coenzyme Q)
- FeS cluster proteins ()
- Cytochromes b, c₁, c, a, a₃ (; )
Nicotinamide Nucleotide-Linked Dehydrogenases
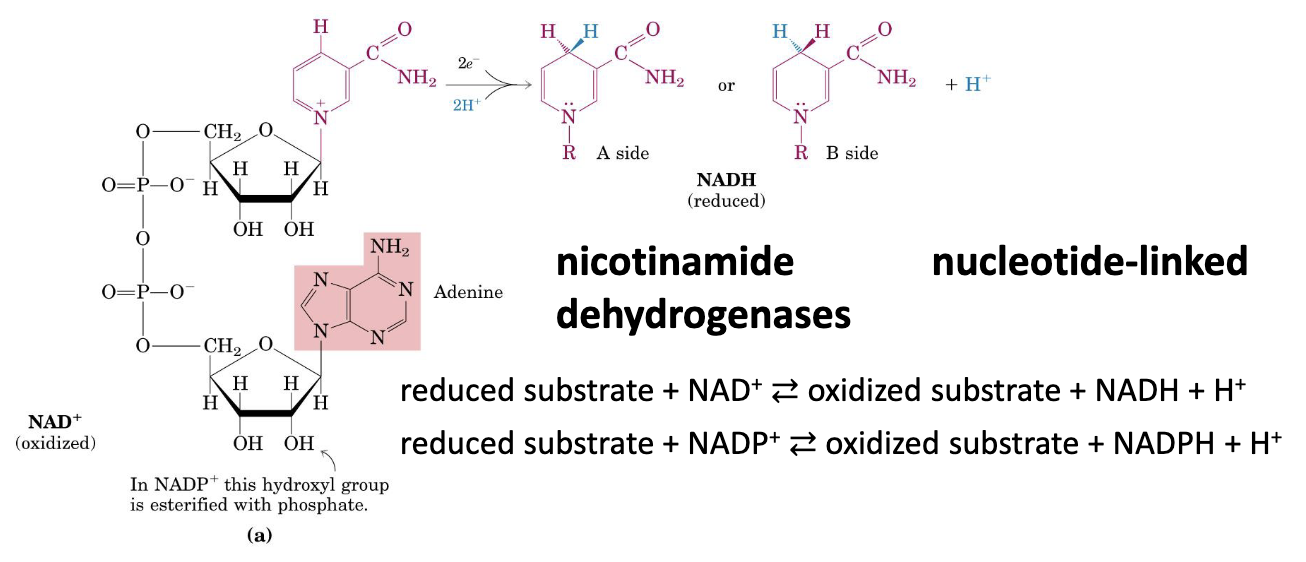
Two hydrogen atoms are removed from the substrates:
- One is transferred as a hydride ion () to
- One is released as in the medium
NADH is not an aromatic molecule anymore. The conjugate something something was broken oh no!!! Since it’s not longer aromatic, it’s unstable.
Why NADH donates electrons readily
NADH transfers a hydride () to the substrate, forming a new C–H bond on the nicotinamide ring. The second proton is released into the medium as . Oxidation back to restores aromaticity, which stabilizes the product and is the thermodynamic driving force for electron donation.
Flavoproteins
Flavoproteins contain a flavin nucleotide (FMN or FAD). The oxidized flavin nucleotide can accept either:
- One (yielding the semiquinone form)
- Two (yielding or )
Unlike NAD⁺,
flavoproteins can accept 1 or 2 electrons, making them flexible intermediaries in the chain.
NAD⁺ vs. Flavoproteins — electron acceptance
accepts only two electrons at once as a hydride (), forming NADH. Flavoproteins (FAD/FMN) can accept one or two electrons, forming either a semiquinone intermediate or the fully reduced form (/). This flexibility makes flavoproteins essential bridge carriers between obligate 2-electron donors and 1-electron carriers like cytochromes.
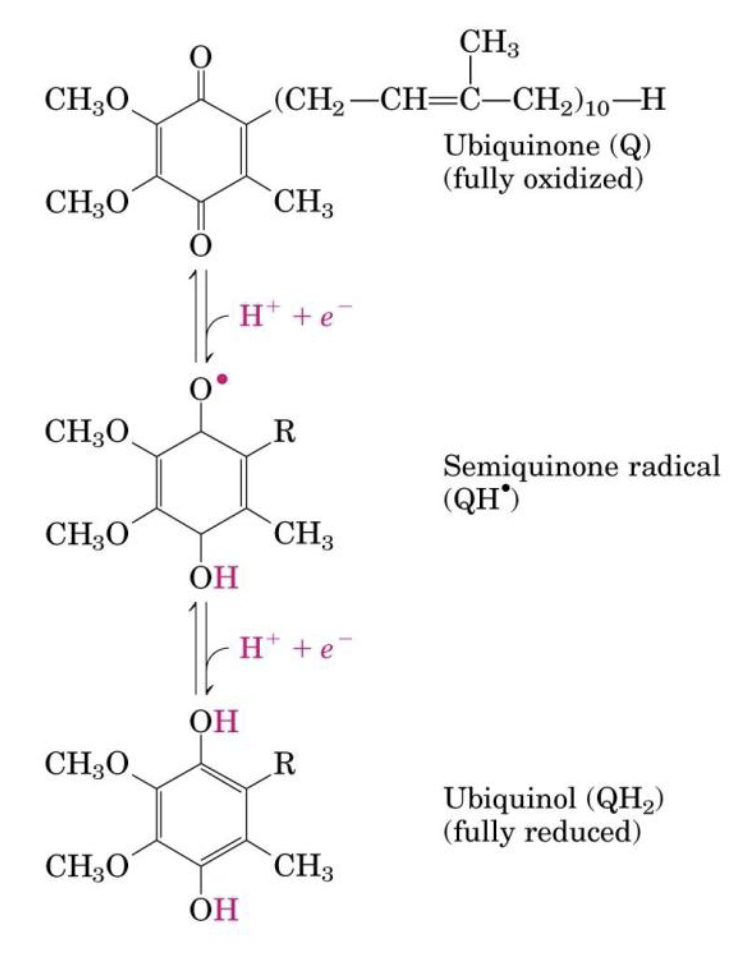
Ubiquinone (Coenzyme Q)
Ubiquinone is a lipid-soluble benzoquinone with a long isoprenoid side chain:
- Can accept one or two
- → there are 3 forms
- Freely diffusible within the inner mitochondrial membrane
- Plays a central role in coupling electron flow to proton movement
CoQ reduction steps
This stepwise reduction is what allows CoQ to bridge 2-electron donors (NADH, FADH₂) and 1-electron carriers (cytochromes).
CoQ is the mobile hub
Its lipid solubility allows it to freely diffuse in the inner membrane, collecting electrons from Complexes I and II and delivering them to Complex III.
Cytochromes
Cytochromes are proteins with iron-containing heme prosthetic groups:
- One carriers ()
- 3 classes in mitochondria: a, b, and c
Heme classes
Class Features Heme b Same heme as in hemoglobin/myoglobin; not covalently bound to the protein Heme c Covalently attached to the protein via Cys residues; found in cytochrome c Heme a Contains a long hydrophobic tail; keeps the cytochrome embedded in the inner mitochondrial membrane
Localization of cytochromes
Cytochromes with hemes a and b are localized within the inner mitochondrial membrane. Cytochrome c is peripheral to the membrane, on the intermembrane space side.
Iron–Sulfur Proteins
Iron–sulfur proteins contain Fe in association with inorganic S atoms and/or the S atoms of Cys residues:
- Participate in one transfers ()
- Sulfur comes from: inorganic sulfide () or Cys residues of the protein
- Rieske iron–sulfur proteins have one Fe atom coordinated to two His residues (instead of two Cys):
- Full coordination: 2 Cys + 2 His
- Found specifically in Complex III (cytochrome complex)
The Rieske Center Logic
The use of His instead of Cys shifts the reduction potential of the Fe-S cluster, allowing it to accept electrons from ubiquinol specifically in the environment of Complex III.
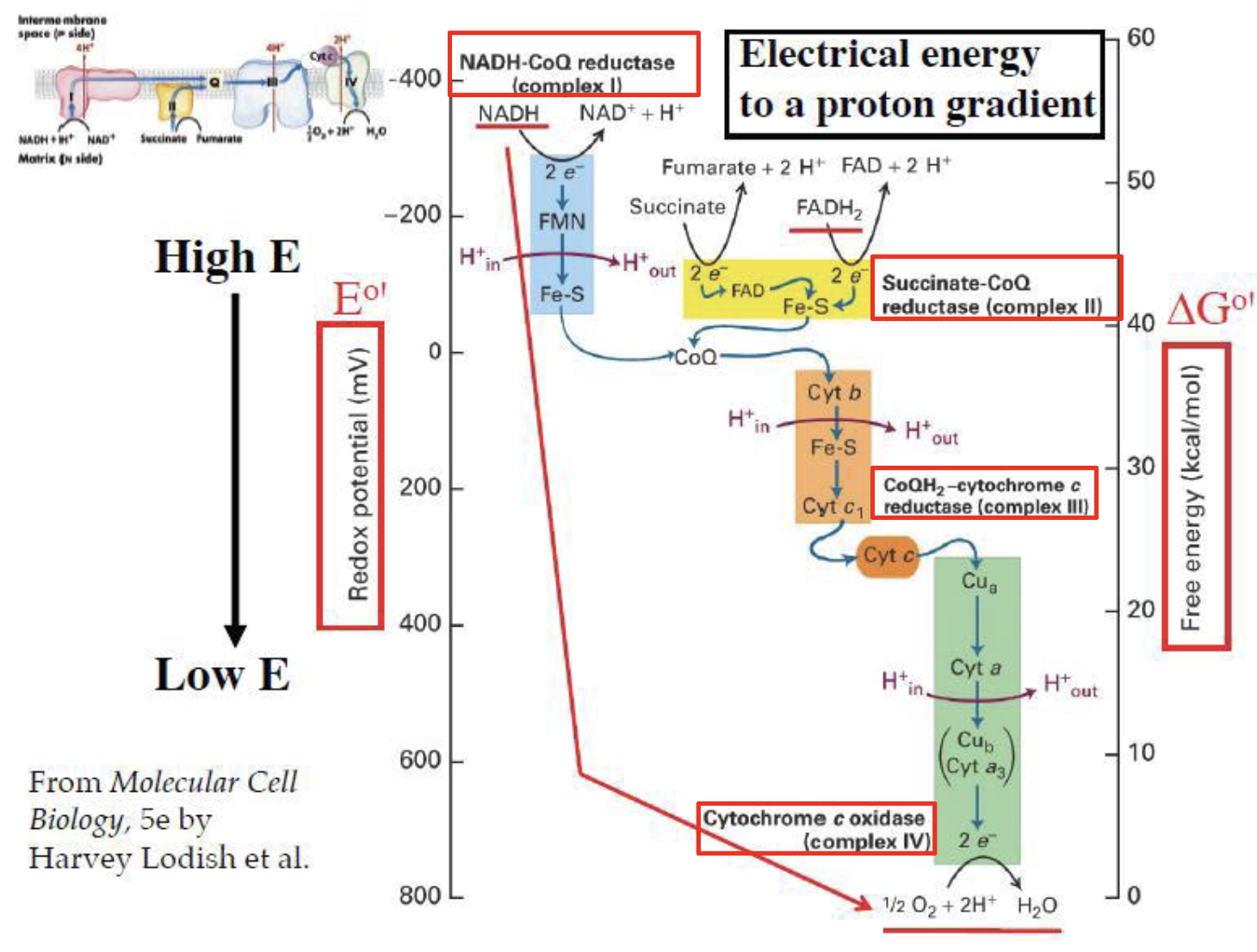
Transfer of electrons from the donor, through a series of proteins, and the final acceptor is oxygen.
This process is spontaneous, it allows the system to use the flow of electrons for a specific chemical process.
The Respiratory Chain — Pathways
Primary pathway:
Why in this direction?
The electron flow is determined by the Standard Reduction Potential ().
- NADH has a very negative potential (), making it a strong electron donor.
- Oxygen has the most positive potential (), making it the ultimate electron acceptor.
- Electrons “fall” down this electrical gradient, releasing free energy ().
Secondary pathway (succinate):
Both pathways converge at CoQ, then share the same route to from Complex III onward.
Proton pumping differs between pathways
- Primary pathway (via NADH → Complex I): 10 H⁺ pumped per electron pair.
- Secondary pathway (via succinate → Complex II): only 6 H⁺ pumped, because Complex II is bypassed and contributes 0 H⁺.
Green demonstrated that these prosthetic groups are contained in individual enzymes called respiratory complexes.
Mitochondrial Respiratory Chain — Four Complexes
(Green and Tzagaloff, 1966): The respiratory chain consists of 4 respiratory multiprotein complexes containing redox coenzymes, connected by mobile intermediates.
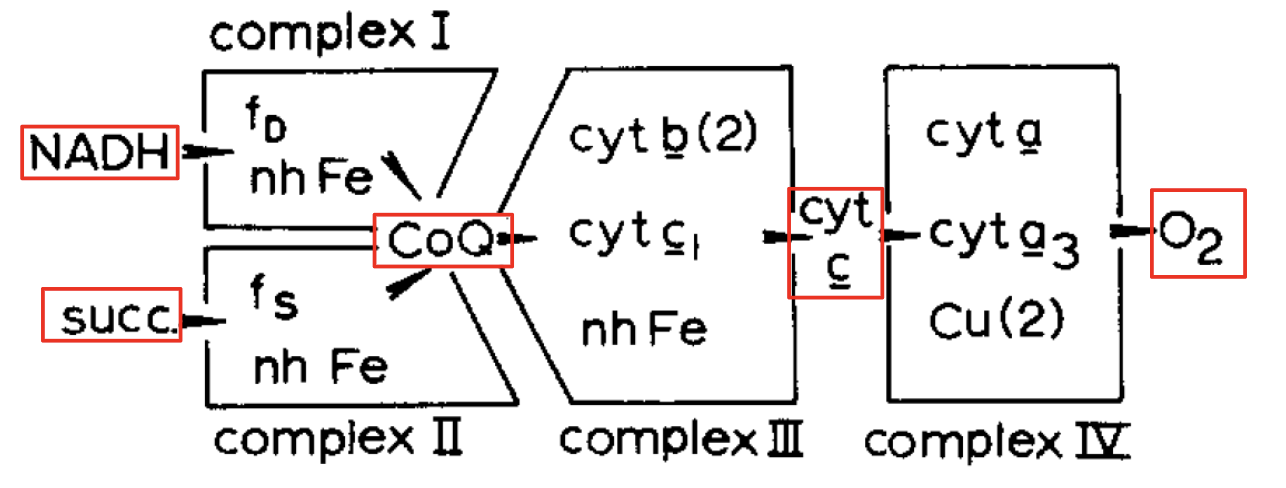
Complex II is unique
Succinate:CoQ reductase (Complex II) is the only membrane-bound enzyme of the TCA cycle. It does not pump protons.
What is unique about Complex II?
Succinate:CoQ reductase is the only membrane-bound enzyme of the TCA cycle, and it does not pump protons.
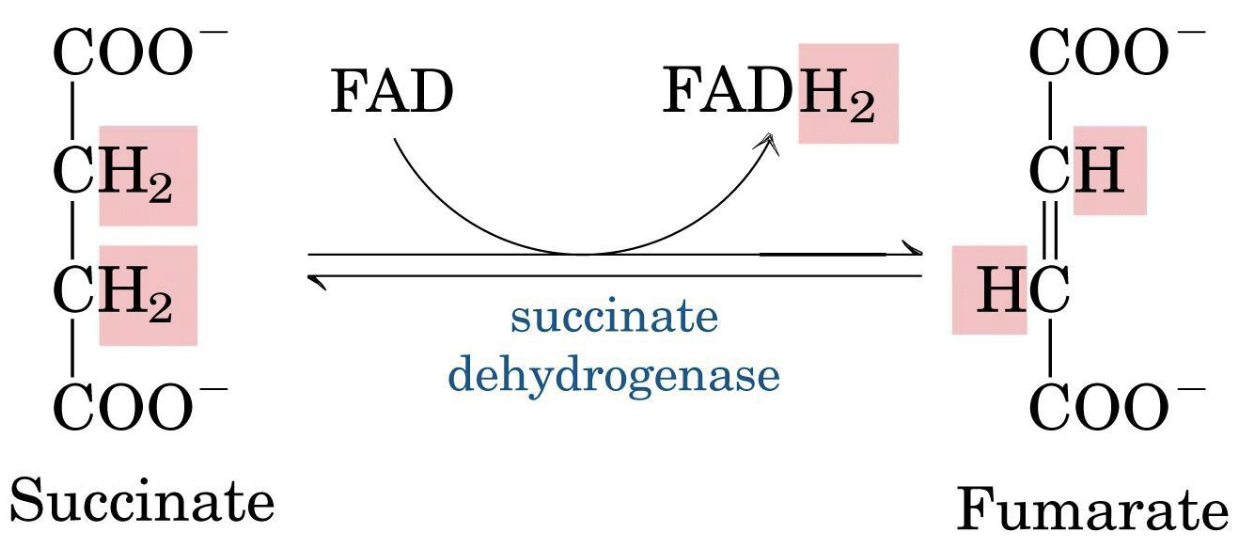
Succinate → Fumarate
This reaction (succinate → fumarate) is the only reaction in common between the respiratory chain and the Krebs cycle.
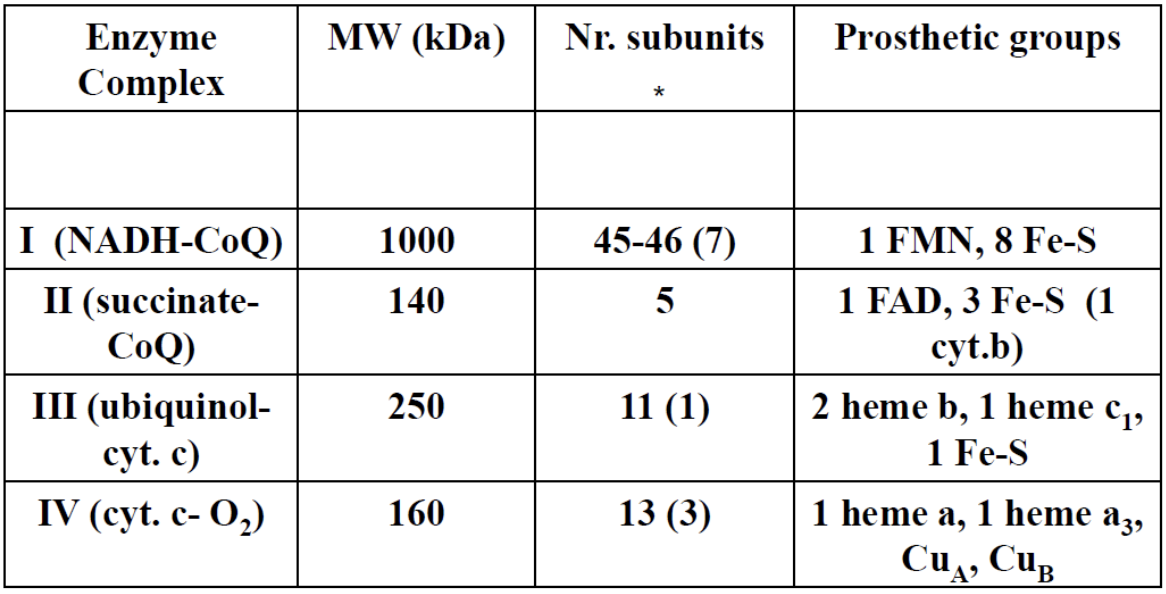
In parenthesis the number of subunits encoded by mitochondrial DNA.
Respiratory Chain Reactions
Complex I — NADH:CoQ Oxidoreductase
It’s performing the redox taking electrons from NADH and giving them to CoQ.
NADH always gives electrons and binds a proton. (?)
It also pumps 4 protons from the mitochondrial matrix to the inter-membrane space (?).
Complex I
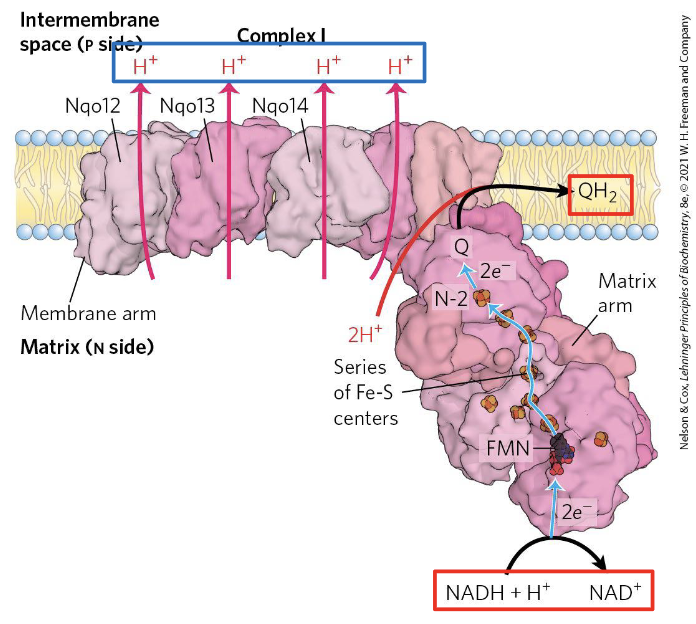
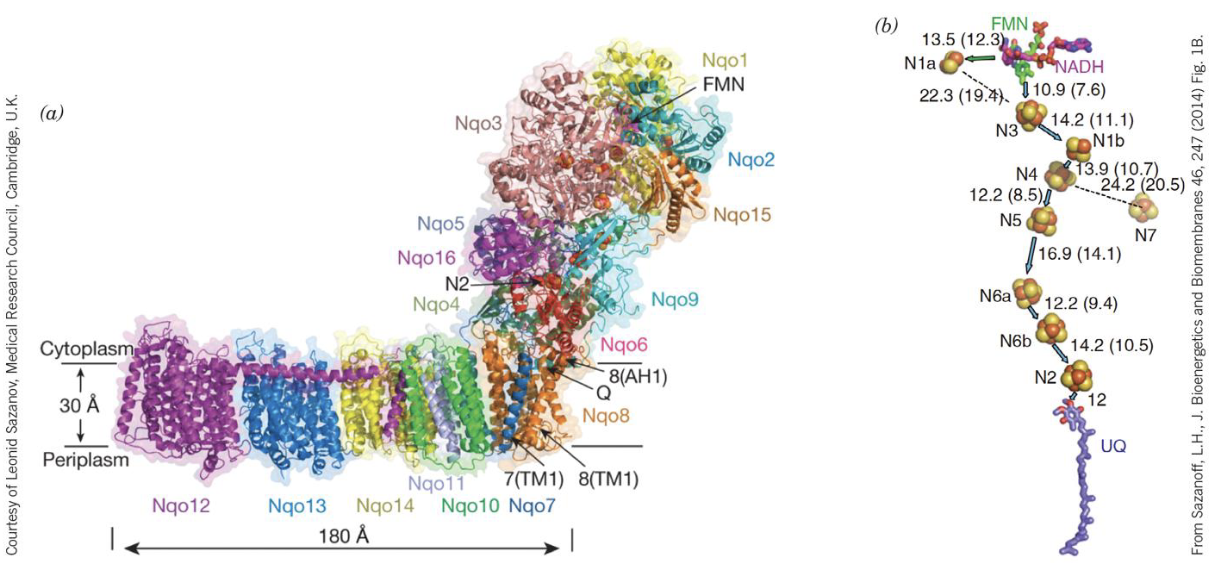
Complex I (NADH:CoQ oxidoreductase) is an L-shaped transmembrane complex. It has two functional arms:
- A hydrophilic arm projecting into the matrix, where NADH is oxidized and electrons enter the chain.
- A hydrophobic arm embedded in the inner membrane, where CoQ is reduced and proton pumping occurs.
Electron flow within Complex I:
NADH donates a hydride () to FMN, the first prosthetic group. Electrons then travel through a series of iron-sulfur (Fe-S) clusters, each passing one electron at a time, until they reach CoQ, which accepts two electrons to become (ubiquinol). This reduction of CoQ is coupled to the pumping of 4 H⁺ from the matrix into the intermembrane space — the mechanism of which involves conformational changes in the membrane arm, though the exact coupling mechanism is still not fully resolved.
Why FMN and not FAD?
Complex I uses FMN (not FAD) as its flavin, because FMN is tightly bound to the complex and can accept the hydride from NADH directly. Its ability to cycle between 1- and 2-electron forms bridges the 2-electron donor (NADH) and the 1-electron Fe-S carriers downstream.
Complex II
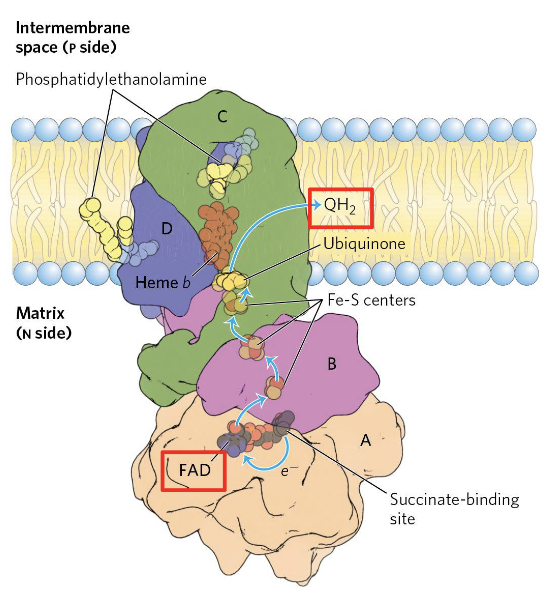
Complex II (Succinate:CoQ reductase, also called succinate dehydrogenase) is the simplest of the four complexes and the only one that does not pump protons. It serves as the direct link between the TCA cycle and the respiratory chain.
Electron flow within Complex II:
Succinate is oxidized to fumarate, and the two electrons are transferred to a covalently bound FAD cofactor, reducing it to . Electrons then pass through Fe-S clusters and reduce CoQ to , which is released into the membrane pool.
Why does Complex II not pump protons?
The redox energy released by the succinate → fumarate reaction ( is small) is insufficient to drive proton translocation across the membrane. All the released energy goes directly into reducing CoQ — none is conserved as a proton gradient. This is why the succinate (FADH₂) pathway yields fewer ATP than the NADH pathway.
Complex II is the TCA–respiratory chain bridge
Succinate dehydrogenase is the only enzyme shared between the TCA cycle and the respiratory chain. It is also the only membrane-bound enzyme of the TCA cycle — all others are soluble in the matrix.
Complex III — CoQH₂:Cyt. c Oxidoreductase
This complex is able to transfer 2 from coq to 2 cyt. c, that contain iron atoms.
We have the transfer of 4 more protons to the inter-membrane space.
Proton count in the Complex III equation
The source material lists the Complex III reaction with consumed, while other formulations (including the equation above) use . The Q-cycle accounts for this: come directly from oxidation (released to IMS), and are drawn from the matrix — giving a net total of 4 H⁺ translocated to the IMS per oxidized.
Internal electron pathway in Complex III
cytochrome Fe–S protein (Rieske center) cytochrome cytochrome
Two cytochrome c molecules are required because each can only carry 1 electron, while donates 2 electrons.
contributes 2 H⁺ directly; 2 more H⁺ are pulled from the matrix → total 4 H⁺ released to the IMS.
Complex IV — Cytochrome c Oxidase
Oxidizing cytochrome c, transferring electrons to oxygen. They are binding to half an oxygen molecule.
Theres a pumping of 2 more protons to the inter-membrane space.
Internal electron pathway in Complex IV
cytochrome cytochromes and + Cu centers
Total
Proton pumping summary
Complex Protons pumped (per NADH) I 4 H⁺ III 4 H⁺ IV 2 H⁺ Total 10 H⁺ Complex II pumps 0 protons.
Thermodynamics of the Respiratory Chain
The standard reduction potential change:
The total potential of this chain:
The standard free-energy change (overall release of energy):
Key thermodynamic values
- per NADH oxidized
Much of this energy is recovered as an electrochemical proton gradient across the inner membrane.
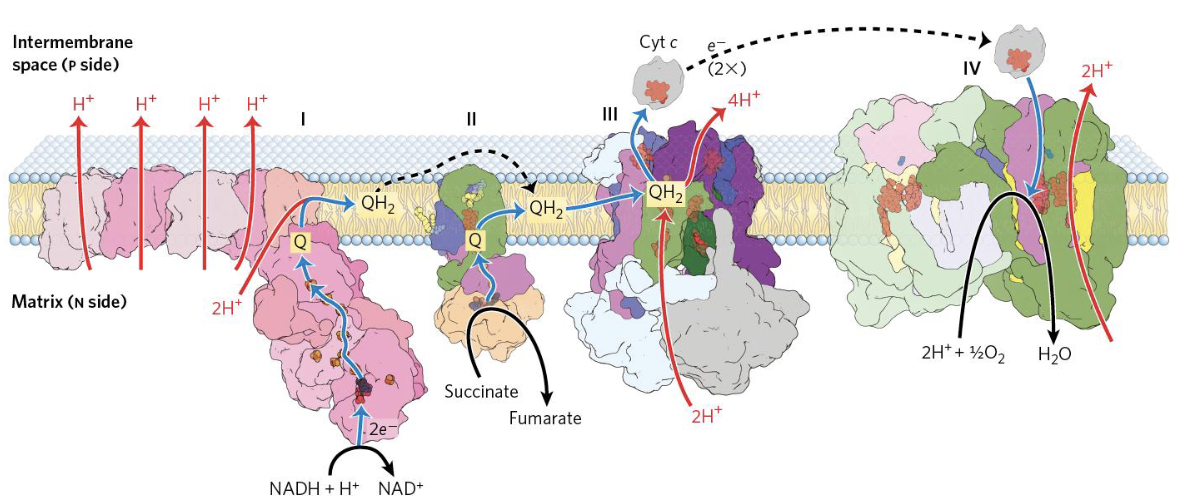
Which side is which?
The matrix is also known as the negative side, because protons are being pumped outside of it.
The intermembrane space instead has a positive net charge,
CoQ has two parts: an aromatic ring that can exchange up to 2 .
The second part is a hydrophobic tail. role? bla bla from complex 1 to complex 3… skipping 2 because that one has a different electron donor: succinate
Cyt3 is on the side of Coq, releasing its only enzyme to ComplexIV. ComplexIII is also pumping outside…
In complexIV two cytC are releasing two electrons to turn two oxygens into water.
not only the entry point of the elctorn donor, but also the net transfer between matrix and intermembrane has to be remembered
todo
Proton-Motive Force
Complexes I, III, and IV are (oxidoreductases but also) proton () pumps toward the exterior, exploiting the energy of redox reactions.
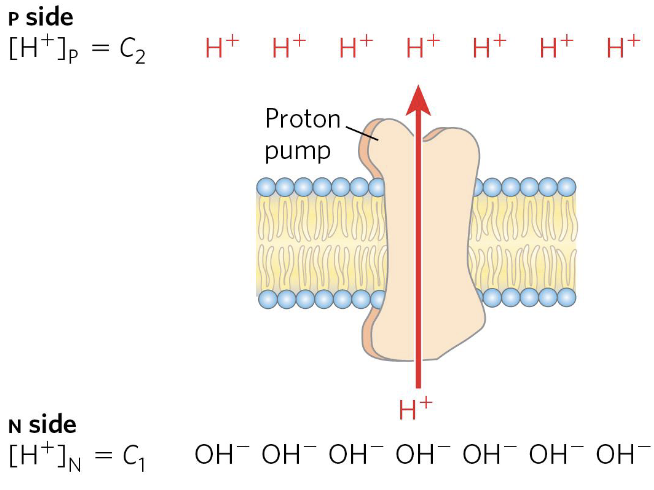
where:
- and are the concentrations of an ion in two regions
- is the absolute value of its electrical charge
- is the transmembrane difference in electrical potential (in volts)
The proton-motive force has two components
- Chemical gradient — across the inner membrane
- Electrical gradient — , the membrane potential
Together they drive ATP synthase (Complex V).
Complex IV → cytochrome oxidase
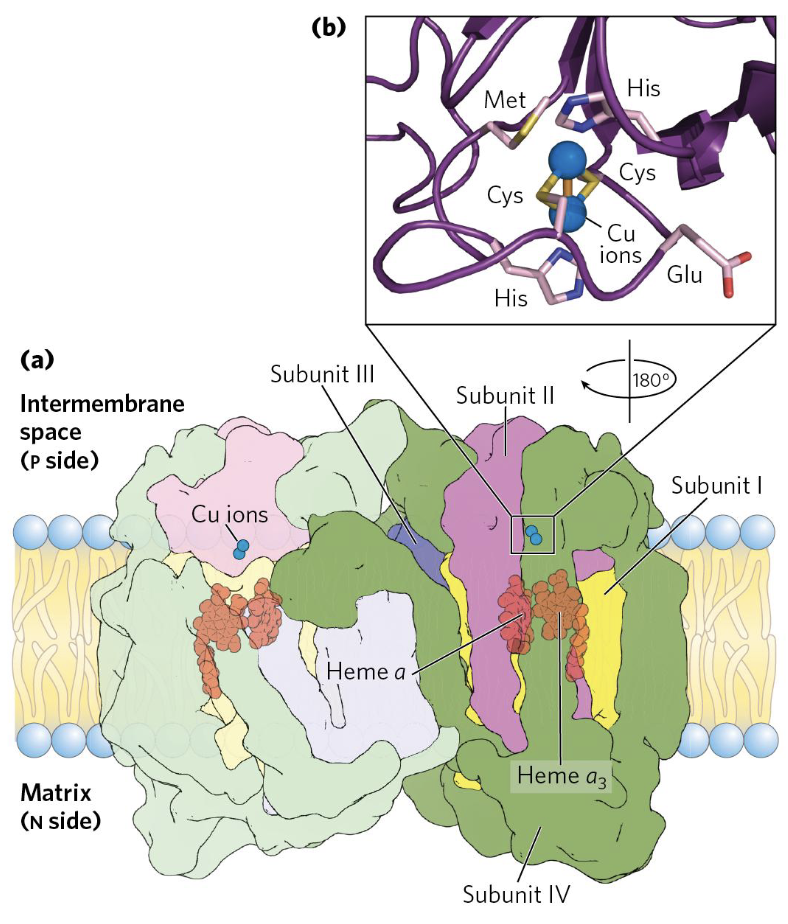
Complex IV (Cytochrome c oxidase) is the terminal enzyme of the respiratory chain. It accepts electrons from cytochrome c (one at a time) and uses them to reduce to — the final, irreversible step.
Electron flow within Complex IV:
- is the initial electron acceptor from cytochrome c.
- Electrons pass to heme a, then to the binuclear center (), where binds and is reduced.
- Four electrons (from four cytochrome c molecules) and four protons from the matrix are needed to fully reduce one to two .
- Simultaneously, 2 H⁺ are pumped into the intermembrane space per electron pair ( reduced).
Why is oxygen the ideal final acceptor?
Oxygen has the most positive standard reduction potential () of any molecule in the chain, making the final reduction step highly exergonic and thermodynamically irreversible. This “pulls” electrons through the entire chain.
Clinical relevance — cyanide poisoning
Cyanide () and carbon monoxide (CO) bind irreversibly to the binuclear center of Complex IV, blocking reduction. This halts the entire respiratory chain, and cells switch to anaerobic metabolism — rapidly fatal in high-dose exposure.
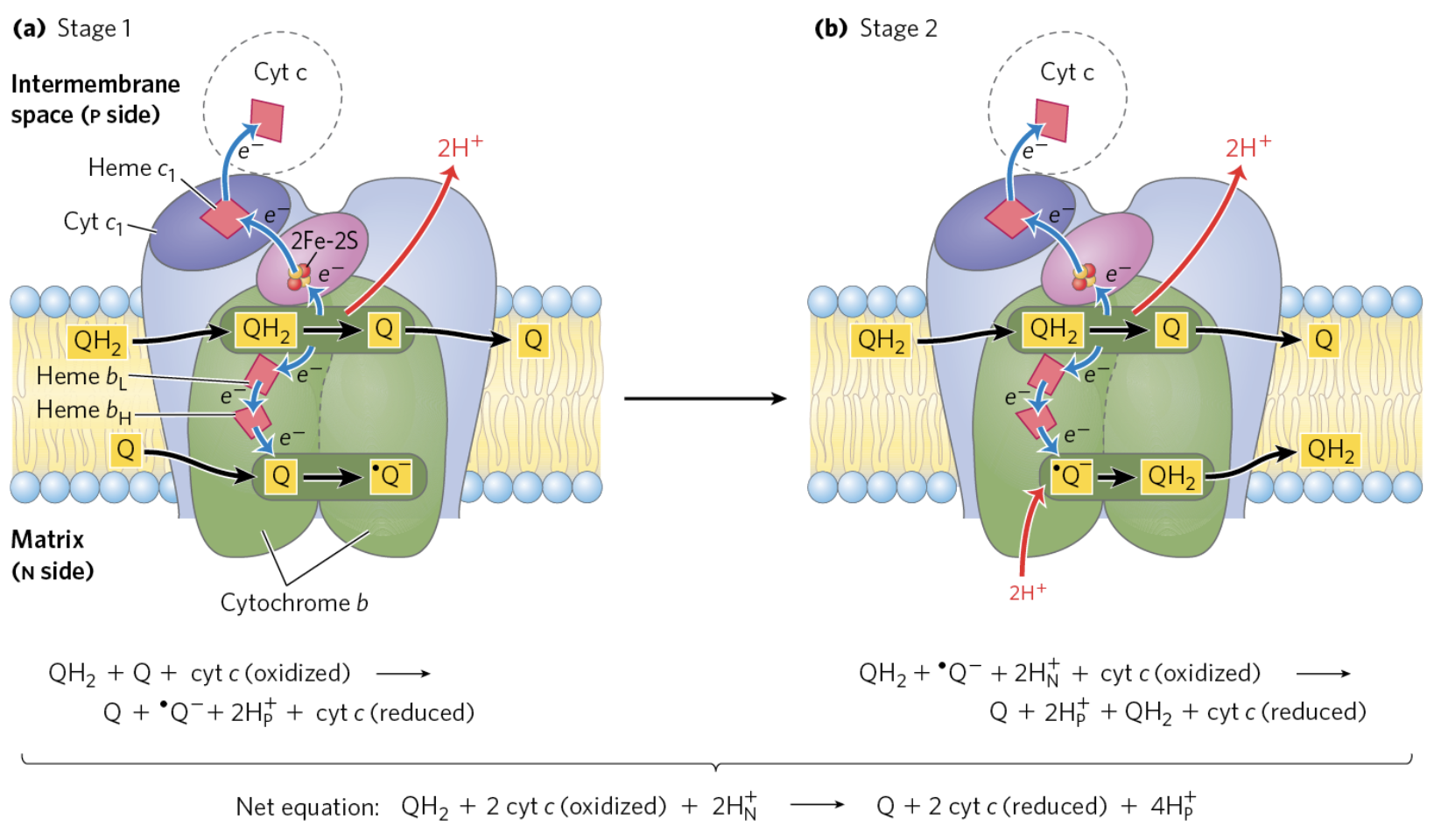
Q-Cycle (Complex III)
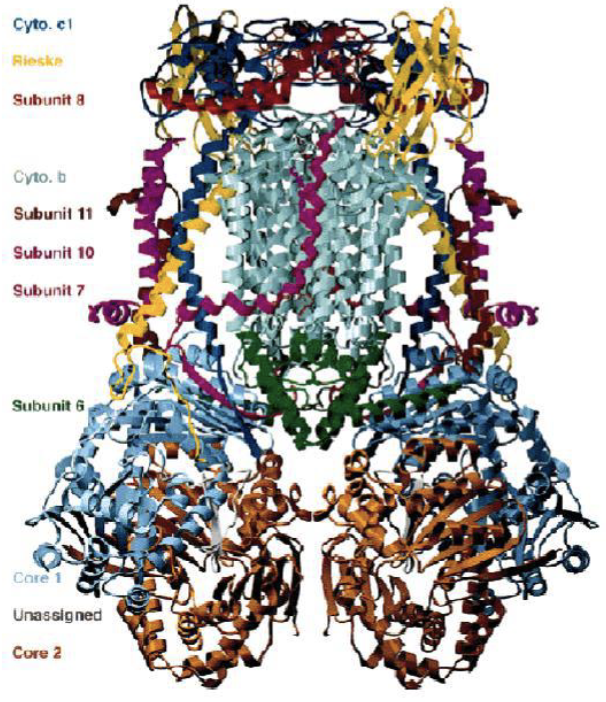 | 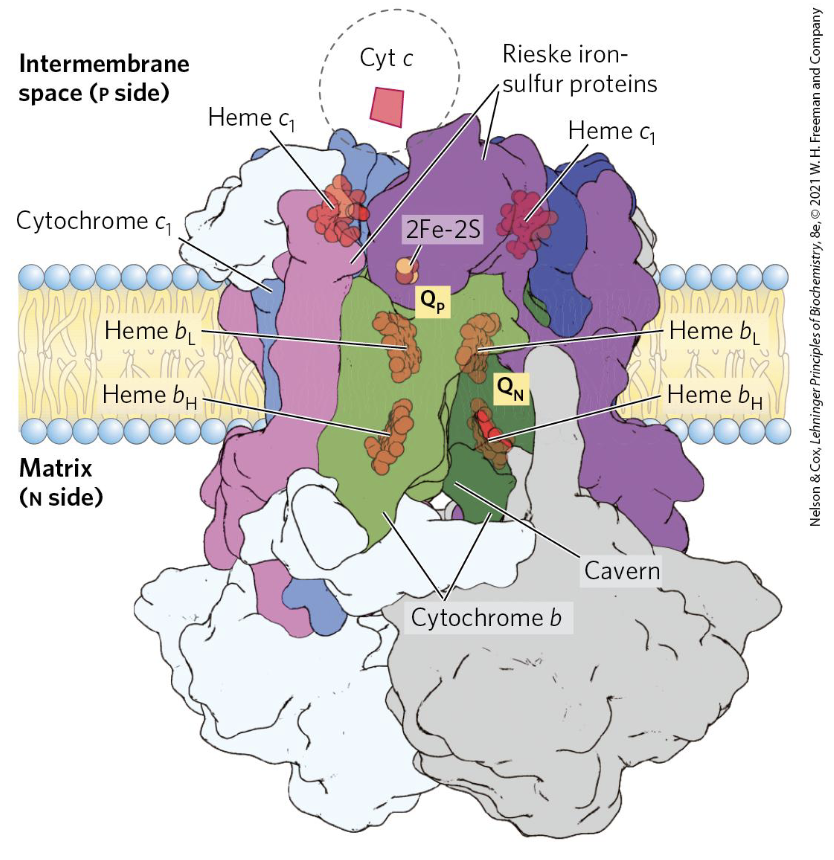 |
|---|
Summary of the Q-cycle:
In stage 1, one of the CoQ (reduced form) enters ComplexIII and has a reaction releasing both electrons that were donated from either Complex I or Complex II.
The second goes to the oxidised CoQ molecule, ubiquinone is reduced only partially into the semiquinone radical.
Total:
The Q-cycle doubles proton pumping
By recycling the semiquinone radical (), Complex III pumps 4 H⁺ per oxidized instead of just 2 — effectively doubling its contribution to the proton gradient.
Three forms of CoQ
Structural Organization of the Respiratory Chain
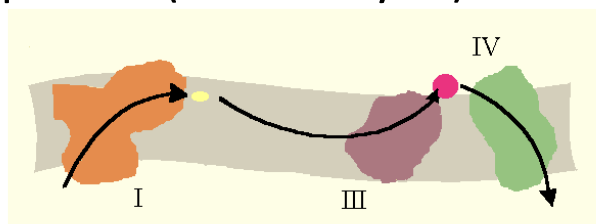
Two models have been proposed:
- Fluid state model: Random collisions between respiratory complexes and mobile components (CoQ and Cyt. c).
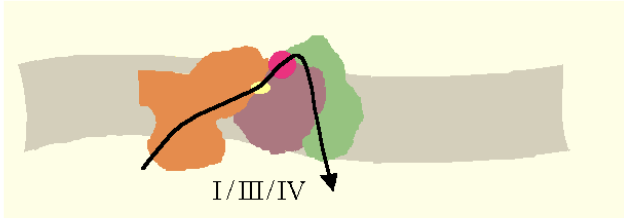
- Solid-state model: Supercomplexes/respirasomes — substrate channelling.
The fluid state model treats the inner membrane as a 2D solution: each complex diffuses independently, and electrons are relayed by random collisions with the mobile carriers CoQ and cytochrome c. While this is thermodynamically plausible, it is kinetically inefficient over long distances.
The solid-state (respirasome) model proposes that Complexes I, III, and IV assemble into stable supercomplexes. Within a respirasome, CoQ and cytochrome c shuttle electrons over very short distances between fixed partners — a process called substrate channeling. Evidence from native gel electrophoresis and cryo-EM supports the existence of these supercomplexes in vivo. Note that Complex II is not part of the respirasome, consistent with its role as an independent TCA-cycle enzyme.
Respirasome
A supercomplex of Complexes I + III + IV in which cytochrome c and ubiquinone diffuse between units. This organization may improve efficiency via substrate channelling, as CoQ does not have to travel long distances.
Why does this matter?
Disruption of supercomplex assembly has been linked to mitochondrial disease and aging. Mutations that destabilize the respirasome reduce the efficiency of oxidative phosphorylation even if individual complexes retain catalytic activity.
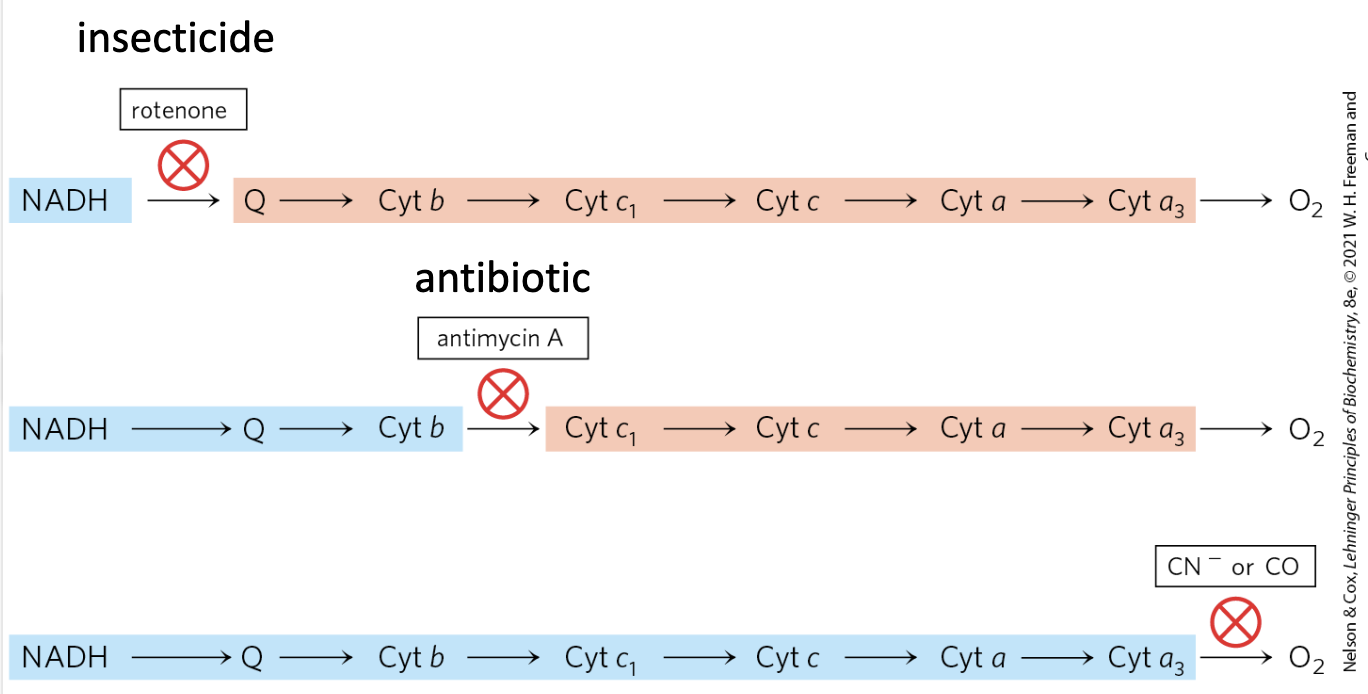
Respiratory Inhibitors
| Inhibitor | Target | Note |
|---|---|---|
| Rotenone | Complex I | Insecticide |
| Antimycin A | Complex III | Blocks electron transfer from Cyt. b to Cyt. c (i.e., at the Qi site) |
| Cyanide / CO | Complex IV | — |
 |
Respiratory inhibitors block the entire chain
Because all complexes are functionally linked, inhibiting any single complex halts electron flow through the entire respiratory chain, shutting down ATP synthesis.
TLDR - 02 - Oxidative Phosphorylation
Detailed Summary
- Core Concept: Oxidative phosphorylation couples the energy released from electron transfer (the Respiratory Chain) to the synthesis of ATP (via a proton gradient).
- Mitochondrial Role: The inner membrane’s impermeability is crucial for maintaining the Proton-Motive Force, which consists of both a chemical () and electrical () gradient.
- Electron Carriers:
- NADH/FADH₂: Primary electron donors.
- Coenzyme Q (Ubiquinone): Mobile, lipid-soluble hub connecting Complexes I/II to III.
- Cytochromes: Proteins with heme groups (Fe) that carry one electron at a time.
- The Respiratory Complexes:
- Complex I: NADH to CoQ; pumps 4 .
- Complex II: Succinate to CoQ; 0 pumped (TCA cycle link).
- Complex III: CoQH₂ to Cyt. c; uses the Q-cycle to pump 4 .
- Complex IV: Cyt. c to ; pumps 2 and forms .
- Thermodynamics: One NADH oxidation yields and pumps a total of 10 .
- Organization: Complexes are likely organized into respirasomes (supercomplexes) to increase efficiency through substrate channeling.
- Inhibition: Blocking one complex (e.g., Rotenone, Antimycin A, Cyanide) halts the entire chain due to functional coupling.
Cytochrome c — Mobile Carrier
Cytochrome c location and function
Cytochrome c is a small, mobile protein located on the outer surface of the inner mitochondrial membrane (facing the IMS).
- Carries 1 electron at a time
- Shuttles electrons from Complex III → Complex IV
- Its heme c is covalently attached via Cys residues (see Cytochromes section above)
ATP Synthase (Complex V)
Role of ATP Synthase
ATP Synthase (Complex V) uses the proton-motive force generated by Complexes I, III, and IV to drive the phosphorylation of ADP → ATP.
The flow of H⁺ back into the matrix through ATP synthase releases the stored electrochemical energy, which is coupled to ATP synthesis.
How many protons are pumped per NADH oxidized through the full respiratory chain?
10 H⁺ total: 4 (Complex I) + 0 (Complex II) + 4 (Complex III) + 2 (Complex IV).
Anki cloze
In the secondary (succinate) pathway, only {1:6} protons are pumped because {1:Complex II} does not pump protons, bypassing the contribution of {1:Complex I}.
Anki cloze
Cytochrome c is located on the {1:outer surface of the inner mitochondrial membrane} and carries {1:one electron} at a time from Complex {1:III} to Complex {1:IV}.
Anki cloze
Rieske iron–sulfur proteins coordinate one iron atom with {1:two histidine residues} (instead of two cysteines) and are found in Complex {1:III}.
Anki cloze
The three heme classes in mitochondria are: heme {1:b} (same as hemoglobin, not covalently bound), heme {1:c} (covalently attached via Cys), and heme {1:a} (has a hydrophobic tail, membrane-embedded).
Anki cloze
Antimycin A inhibits Complex {1:III} by blocking electron transfer from cytochrome {1:b} to cytochrome {1:c}.
Anki cloze
Cytochromes with hemes a and b are located {1:within} the inner mitochondrial membrane, while cytochrome c is {1:peripheral to it, on the intermembrane space side}.