TARGET DECK: MED::I::Signaling Pathways in Health and Disease::Metabolic Biochemistry::03 - Oxidative Phosphorylation P2
Chemiosmotic Coupling
The chemiosmotic model describes how ATP synthesis is coupled to an electrochemical proton gradient, known as the proton-motive force (PMF):
- The respiratory chain pumps H⁺ from the matrix → intermembrane space (IMS), exploiting the negative of electron transfer.
- The resulting proton gradient stores potential energy.
- Protons passively return to the matrix through ATP synthase, driving ATP synthesis.
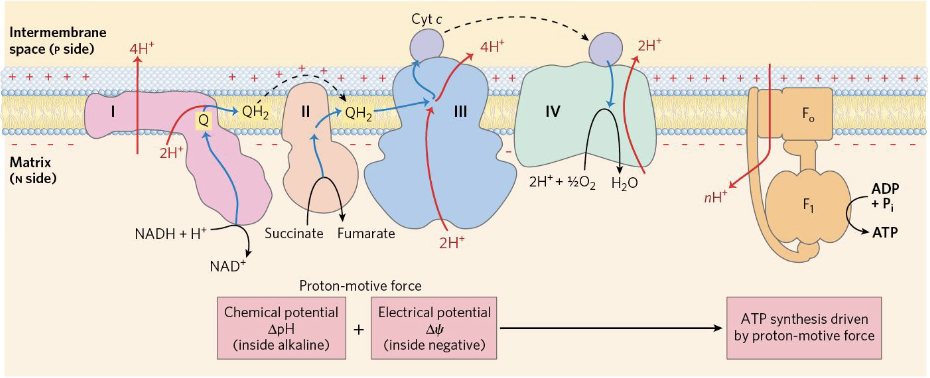
The equation for ATP synthesis is:
Key conceptual shift
Classical biochemistry assumed that coupling an endoergonic reaction to an exoergonic one required a chemical intermediate. The chemiosmotic hypothesis replaced this with a physical intermediate — the proton gradient itself.
What does the chemiosmotic model propose?
ATP synthesis is coupled to an electrochemical proton gradient, and proton flow back through ATP synthase drives ATP production.
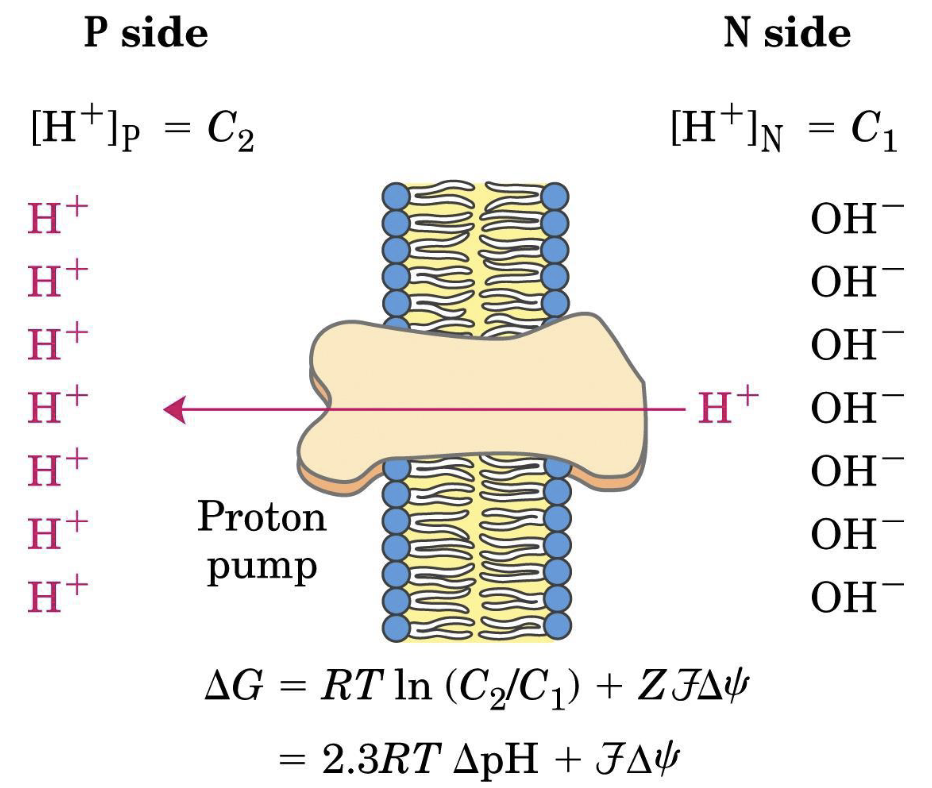
This is the role of complex I, III, IV.
Organization of the Respiratory Chain: Two Models
Two models have been proposed for how the respiratory complexes are organized in the inner mitochondrial membrane:
- Fluid-state model: Complexes diffuse randomly; electron transfer depends on random collisions between complexes and the mobile carriers CoQ and Cyt. c.
- Solid-state model (supercomplexes/respirasomes): Complexes associate into stable supercomplexes. The respirasome contains Complexes I, III, and IV together; CoQ and Cyt. c diffuse within the supercomplex, enabling substrate channelling. This model is currently more accepted.
The {{respirasome}} is a supercomplex containing Complexes {{I, III, and IV}}, in which {{cytochrome c and ubiquinone}} diffuse between subcomplexes to enable substrate channelling.
Stoichiometry of Proton Consumption
Protons re-enter the matrix through ATP synthase, obligatorily coupled to ATP synthesis ().
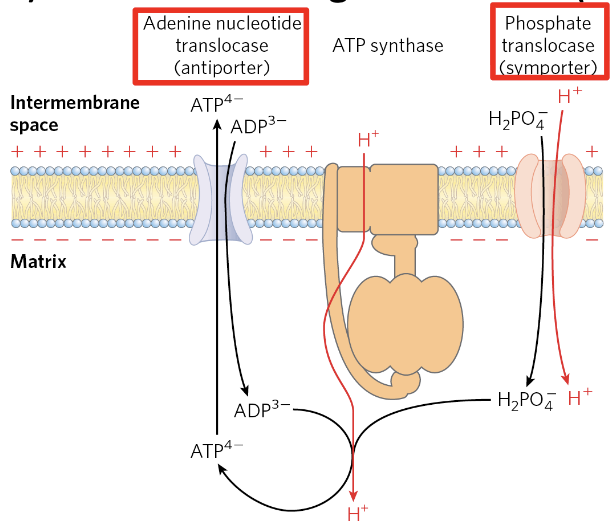
However, ADP and must also be imported from the cytosol, consuming an additional :
- Phosphate import: symport with (electroneutral, since is negatively charged).
- ATP/ADP exchange: ATP (4⁻) exits in exchange for ADP (3⁻) → net export of 1 negative charge (electrogenic).
Total cost
(ATP synthase) (transport) per ATP synthesized.
How many protons are required per ATP synthesized in oxidative phosphorylation?
4 H⁺ per ATP: 3 H⁺ through ATP synthase plus 1 H⁺ for transport.
Stoichiometry of Oxidative Phosphorylation
NADH oxidation (Complex I entry)
- pumped out per
- required per ATP
Succinate oxidation (Complex II entry)
- pumped out per
Scalar vs. vectorial protons
At Complex I, the oxidation releases in the matrix. These are scalar (not vectorial) protons — they are recovered during CoQ → CoQH₂ reduction, still on the matrix side, so they do not affect the pumping stoichiometry.
ATP Synthase (F₁F₀-ATPase)
Double nature
This enzyme can both hydrolyze ATP and synthesize ATP.
- The hydrolysis happens if the subunit is isolated,
- the synthesis happens if the subunits are together and coupled to a proton gradient.
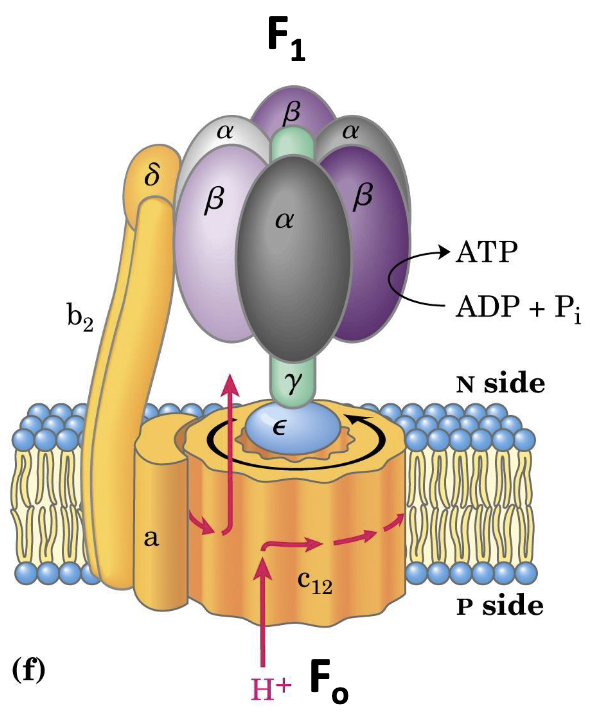
Subunit architecture:
| Component | Location | Subunits | Function |
|---|---|---|---|
| F₀ | Integral membrane | Proton channel; – interface forms the pore - (hydrophobic) | |
| F₁ | Peripheral (matrix) | Catalytic head; subunits carry the 3 active sites - (hydrophilic) |
The subunit of has a variable number. In homo sapiens, is .
- Subunit is immobilized by contact with subunit , which also anchors .
- Proton entry from IMS drives rotation of the -ring.
- -ring rotation drives rotation of the subunit within , cyclically exposing active sites.
The actual synthesis happens in the subunits of the component.
What are the roles of the F₀ and F₁ parts of ATP synthase?
F₀ is the membrane proton channel, and F₁ is the catalytic head that synthesizes ATP.
Reverse mode: ATP synthase as ATPase
If the proton gradient is absent or insufficient, the enzyme reverses: it hydrolyses ATP to pump protons toward the IMS, thereby generating a proton gradient. In this mode it functions as an ATPase. The enzyme cannot catalyse only part of the overall reaction — it requires either the gradient (for synthesis) or ATP (for hydrolysis).
Anki cloze
In the absence of a proton gradient, ATP synthase {{hydrolyses ATP}} to {{pump protons into the IMS}}, generating a proton gradient at the cost of ATP.
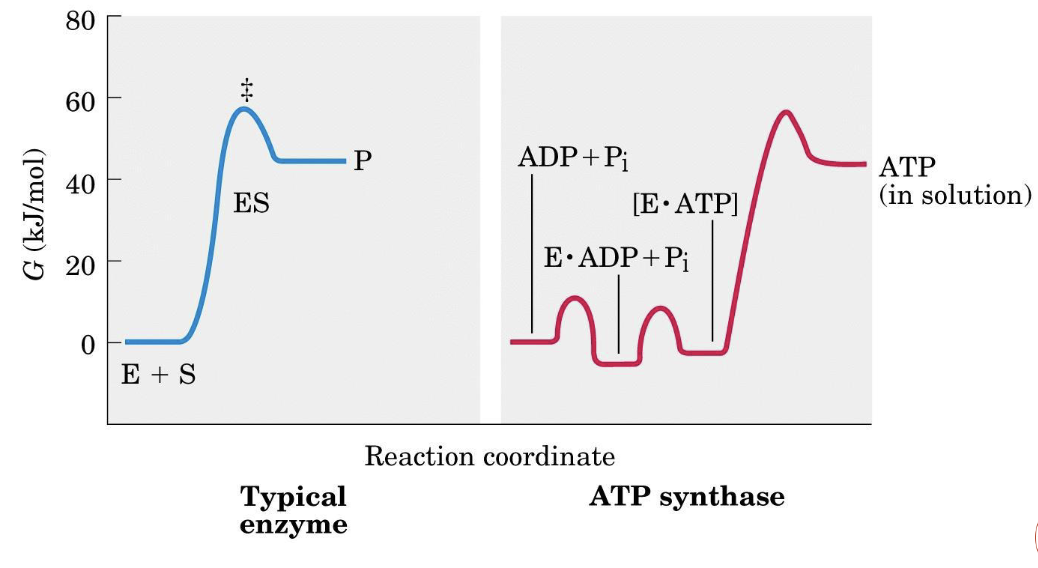
Why ATP Synthesis Needs No Energy in the Active Site
In aqueous solution:
The standard state includes . In the hydrophobic active site of ATP synthase, water is excluded:
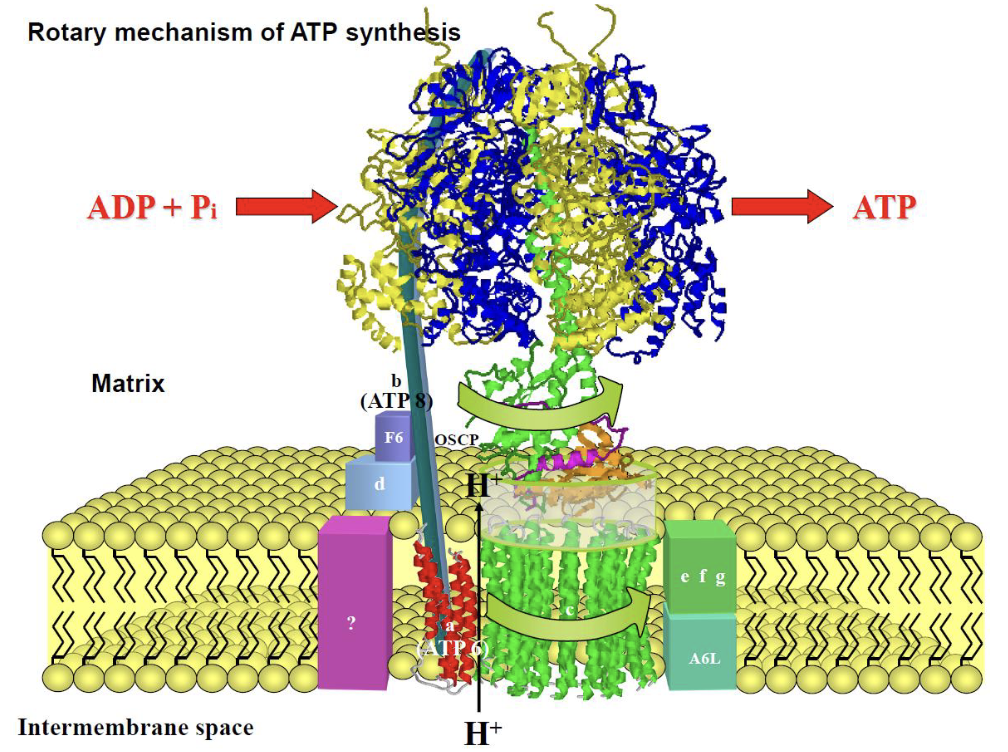
Binding-change mechanism
ATP synthesis in the active site is essentially thermodynamically spontaneous. The energy from the proton-motive force is used to release ATP into the aqueous medium via a conformational change — not to form the bond itself.
Binding-Change Model
Rotational catalysis
Is the mechanism by which the flow of protons through causes the ring to rotate and, in turn, trigger the subunit conformational changes in .
Each subunit cycles through three conformational states:
-
-ATP — tight binding (product held)
-
-ADP — loose binding
-
-empty — very loose binding (open)
-
translocated → release of ATP
-
full rotation → synthesis of ATP
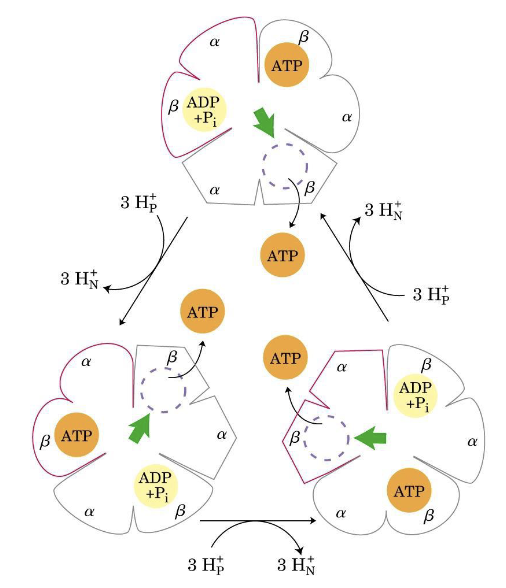
Subunit roles in rotational catalysis
- α subunits: structural/regulatory — do not catalyse ATP synthesis.
- β subunits (×3): each carries an active site; they cycle through L → T → O conformations.
- γ subunit: the rotating “shaft” or engine; each 120° rotation changes one β subunit’s conformation.
- Every 3 H⁺ translocated through F₀ → one β subunit completes a conformation cycle → release of 1 ATP.
Respiratory Control
Control mechanism
Protons can only traverse ATP synthase coupled to ATP synthesis — if ADP is absent or ATP synthase is inhibited (e.g. by oligomycin), proton flow stops.
When Reaction 1 reaches electrochemical equilibrium, respiration ceases. The residual rate reflects passive H⁺ leak across the inner membrane; membrane damage increases this leak.
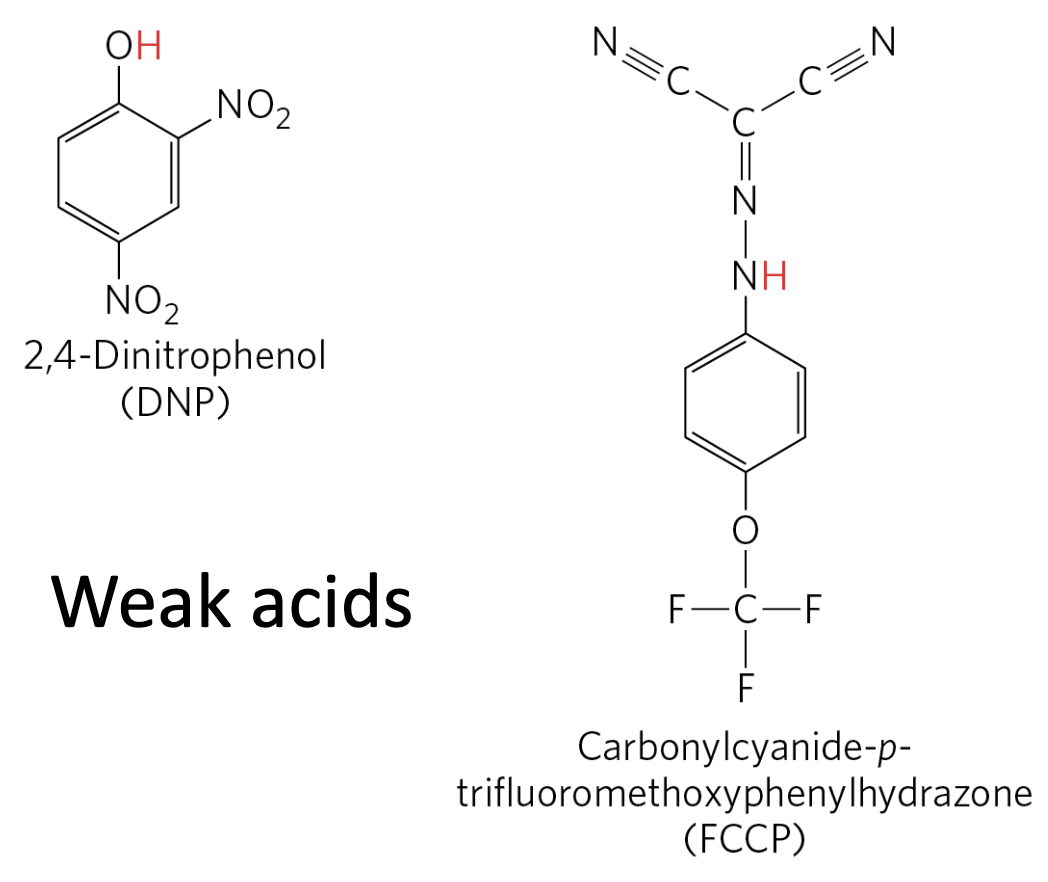
Uncouplers
Uncouplers are membrane-permeant weak acids that carry protons across the inner mitochondrial membrane (back to the matrix), dissipating the proton gradient without passing through ATP synthase.
- ✗ ATP synthesis is blocked
- ✓ Electron transport continues (or accelerates)
What do uncouplers do?
They dissipate the proton gradient across the inner mitochondrial membrane, blocking ATP synthesis while allowing electron transport to continue.
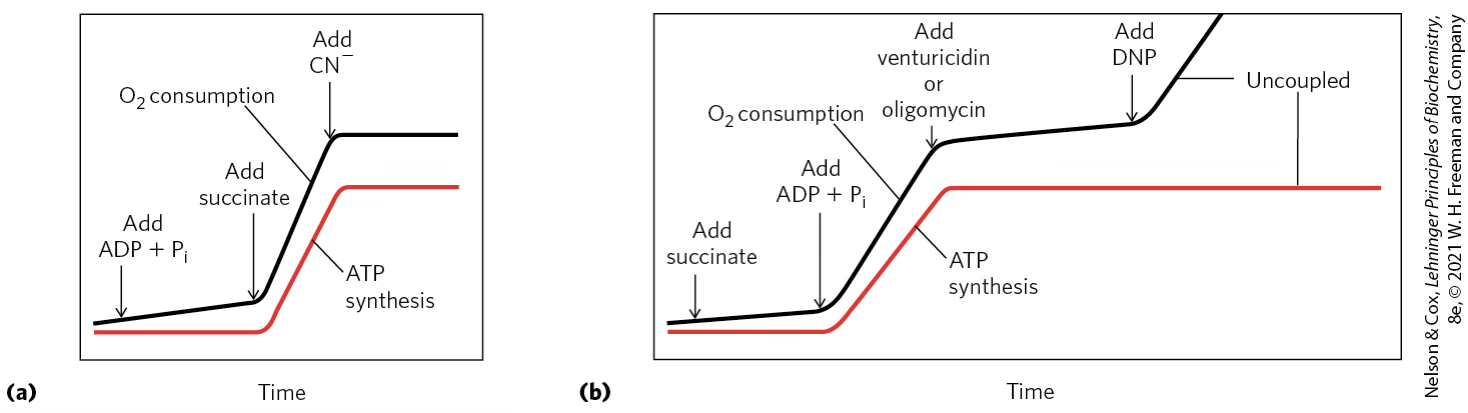
Classic experimental sequence with succinate
- Succinate alone → ET builds up until equilibrium; O₂ consumption slows.
- + ADP + → protons flow through ATP synthase; ET resumes; O₂ is consumed.
- + Oligomycin → ATP synthase blocked; no proton flow; ET again rebuilds gradient.
- + DNP (uncoupler) → gradient dissipated; ET proceeds without ATP synthesis. The protons are going back to the matrix without passing through ATP synthase.
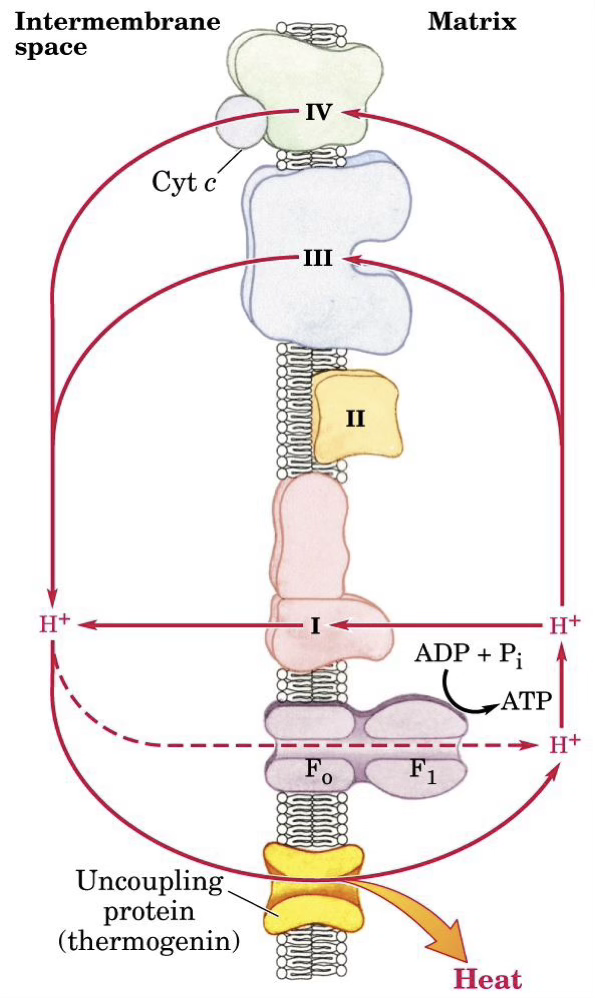
Thermogenin
Protons that go back inside the membrane do so spontaneously. Thermogenin is an uncoupler that allows this process to happen resulting in the production of heat.
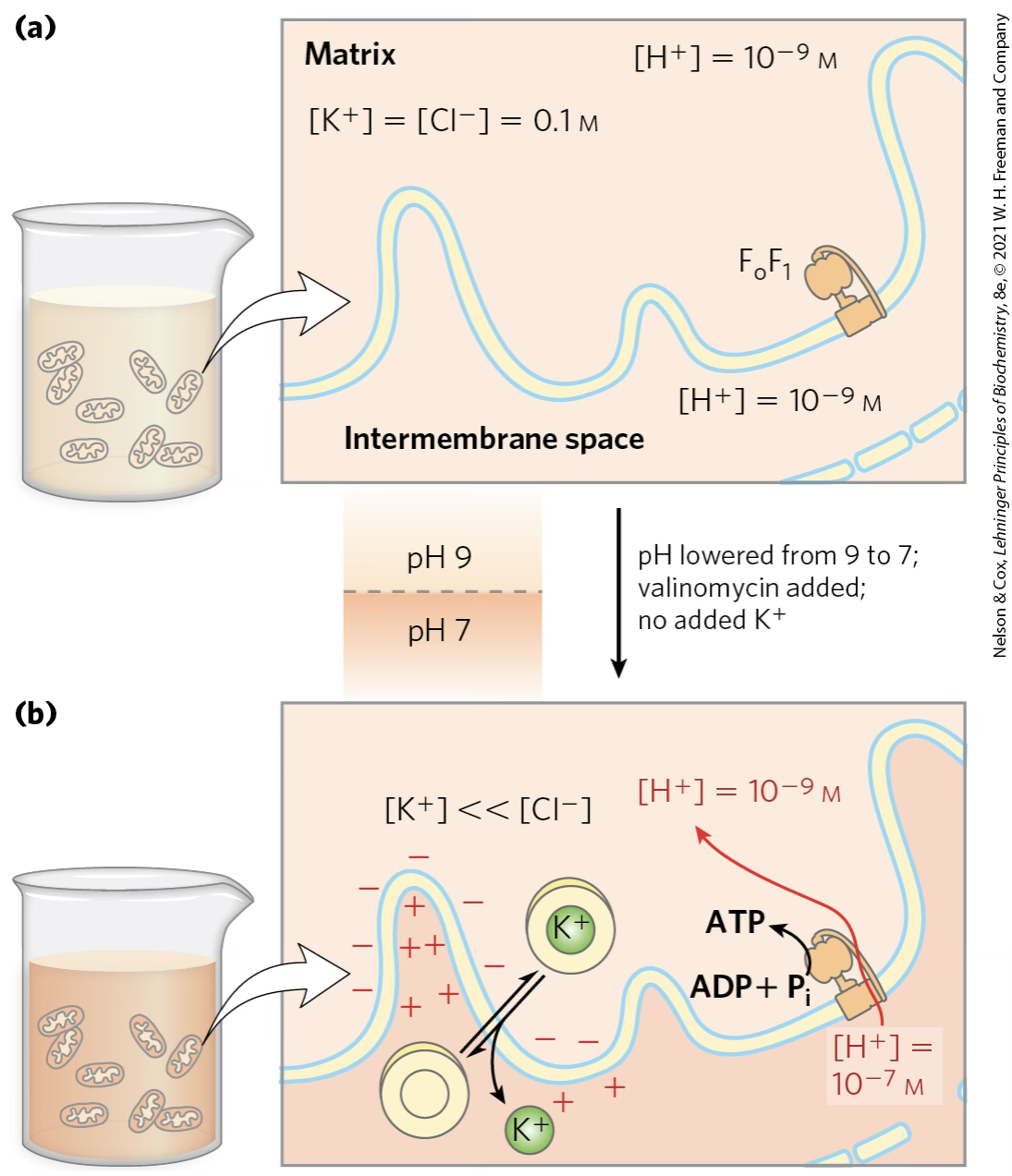
Valinomycin
In the absence of an oxidizable substrate, the proton-motive force alone drives ATP synthesis without electron transport.
Valinomycin is a polypeptide used for K transport; it allows K⁺ to flow out of the matrix. As positive charges (K⁺) leave the matrix, the inside becomes electrically negative relative to the outside. No extra K⁺ is added to the medium, so K⁺ keeps leaving → [K⁺] << [Cl⁻] outside.The result is that both components of the proton-motive force are now in place:
- A chemical gradient (ΔpH): more H⁺ outside than inside;
- An electrical gradient (Δψ): inside negative, outside positive.
These two gradients together drive H⁺ through Fo and into the matrix — and the FoF₁ ATP synthase converts ADP + Pᵢ into ATP, even though the electron transport chain never operated at any point.
→ It is possibl eto make ATP synthase work even if the electron transfer system no longer functions.
→ This experiment, historically important for validating chemiosmotic theory, proves that the proton-motive force is the direct and sole energy input needed by ATP synthase — it doesn’t matter how the gradient is generated (respiration, artificial acidification, or anything else). The enzyme simply harnesses the flow of protons down their electrochemical gradient.
OxPhos Inhibitors Summary
| Class | Example | Primary Target | Effect on ET | Effect on ATP synthesis |
|---|---|---|---|---|
| Respiratory inhibitors | Rotenone, Antimycin A, KCN | Respiratory chain | Blocked | Blocked (secondary) |
| ATP synthase inhibitors | Oligomycin | Slows | Blocked | |
| Uncouplers | DNP, FCCP | Inner membrane | Increases | Blocked |
Membrane Potential: Other Uses
The across the inner membrane also drives transport:
- Ca²⁺ uniport — calcium influx (inward)
- Phosphate/ cotransport — symport (inward)
- ADP³⁻/ATP⁴⁻ exchange — electrogenic (net exit of negative charge)
- Pyruvate/ exchange — net exit of negative charge
Mitochondria and Pathology
Mitochondria have their own mtDNA.
Reactive Oxygen Species (ROS)
Incomplete reduction of generates reactive intermediates:
Sources of ROS:
- Ionizing/UV radiation, xenobiotics, smoking
- Transition/heavy metal ions
- NADPH oxidase (phagocytes)
- Microsomal oxidations (cytochrome P-450)
- Amine oxidases, xanthine oxidase
- Mitochondrial respiratory chain (major endogenous source)
ROS also participate in intracellular signalling — partly through oxidative inactivation of phosphatases.
Mechanisms of protection:
- Induction of protective factors: NF-κB, HSP
- Repair mechanisms
- Removal: proteolysis, autophagy, apoptosis
- Antioxidant enzymes: SOD, GPx, catalase
- Chemical antioxidants (water- and lipid-soluble)
Mitochondrial Theory of Aging and Disease
Central thesis
Most human pathologies share a common mitochondrial mechanism: ROS overproduction + accumulation of somatic mtDNA mutations → loss of OXPHOS capacity → cellular energy failure and death.
Molecular cascade:
The vicious cycle: mtDNA mutations impair the respiratory chain → more electron leak → more ROS → more mtDNA damage → …
Pathologies linked to this mechanism (aging as the major predisposing factor):
- Ischemic diseases (myocardial infarction, stroke)
- Degenerative skeletal diseases
- Neurodegenerative diseases (Parkinson’s, Alzheimer’s)
- Type 2 diabetes
- Cancer
Historical note
The discovery in the 1980s that mtDNA mutations cause hereditary diseases launched the field of mitochondrial cytopathology. Respiration/OXPHOS defects increase glycolysis → lactic acidosis, and often increase ROS production.
TLDR - 03 - Oxidative Phosphorylation
Chemiosmotic coupling links the respiratory chain to ATP synthesis via a proton gradient (proton-motive force). The chain pumps from the matrix to the IMS; protons flow back through ATP synthase, driving ATP production. The true energy cost is ( through + for metabolite transport). This gives a P/O ratio of 2.5 for NADH oxidation and 1.5 for succinate oxidation.
ATP synthase () works by rotational catalysis: protons spin the -ring → rotating subunit cycles subunits through three conformations (tight/loose/open). ATP bond formation itself costs almost nothing in the hydrophobic site; the energy is spent releasing ATP into solution. Each full rotation produces 3 ATP, consuming .
Respiratory control tightly couples electron transport to ATP demand — if ADP is absent or ATP synthase is inhibited (oligomycin), respiration stops. Uncouplers (DNP, FCCP) are weak acid proton carriers that short-circuit the gradient, allowing electron transport to run unchecked while blocking ATP synthesis — effectively converting the PMF to heat.
ROS — superoxide, , and the hydroxyl radical — are generated mainly by the mitochondrial respiratory chain. They damage mtDNA (which is unprotected and close to the chain), impairing OXPHOS and generating yet more ROS. This vicious cycle is central to the mitochondrial theory of aging: progressive mtDNA mutation accumulation → energy failure → cell death — underlying ischemic disease, neurodegeneration, diabetes, and cancer.