TARGET DECK: MED::I::Morphology and Development::Histology::10 - Muscle Tissue
Learning Objectives
- Identify the three types of muscle at the light microscope level, including distinctive features of each (e.g., intercalated disk of cardiac muscle).
- Describe the structural basis of muscle striation.
- Know the structural elements that harness muscle contraction (shortening of myofibrils) to the movement of a body part, and the mechanism by which muscle cells contract.
- Understand the function and organization of connective tissue in skeletal muscle (endo-, peri-, and epimysium).
- Be familiar with the regenerative potential of each muscle type.
Overview: Types of Muscle Tissue
| Type | Striation | Control | Location |
|---|---|---|---|
| Skeletal | Striated | Voluntary | Locomotion, mastication, phonation |
| Cardiac | Striated | Involuntary | Heart, pulmonary veins |
| Smooth | Non-striated | Involuntary | Blood vessels, viscera, dermis |
Embryonic origin
All muscle tissues develop from the mesoderm, including cardiac muscle, skeletal muscle, and smooth muscle of the gut.
Anki cloze
Muscle tissues develop from the {1:mesoderm}.
Properties of Muscle Tissue
| Property | Definition |
|---|---|
| Excitability | Responds to chemicals from nerve cells; generates electrical signals |
| Contractility | Ability to shorten and generate force |
| Extensibility | Ability to stretch |
| Elasticity | Ability to return to original shape |
Functions of Muscle Tissue
- Produce movement
- Maintain posture
- Stabilize joints
- Thermogenesis
Cachexia = loss of weight, muscle atrophy, fatigue, weakness.
Muscle-Specific Terminology
| General term | Muscle-specific term |
|---|---|
| Cell membrane | Sarcolemma |
| Cytoplasm | Sarcoplasm |
| Smooth endoplasmic reticulum | Sarcoplasmic reticulum |
| Mitochondria | Sarcosomes |
Anki cloze
In muscle cells, the smooth endoplasmic reticulum is called the {1:sarcoplasmic reticulum}, and the cell membrane is called the {2:sarcolemma}.
Skeletal Muscle
General Characteristics
- Long, cylindrical, multinucleated cells
- Nuclei located peripherally (just beneath the sarcolemma)
- Contraction: quick, vigorous, voluntary
- Size: few mm up to 10 cm in length; 10–100 µm in width
Sarcoplasm Contents
- Several mitochondria
- Well-developed endoplasmic reticulum
- Glycogen granules
- Myoglobin
- Contractile proteins organized into myofibrils (~1–3 µm diameter → provide striation)
Anki cloze
Skeletal muscle fibers are {1:multinucleated} cells with nuclei located {2:peripherally}, just beneath the sarcolemma.
Connective Tissue Investments of Skeletal Muscle
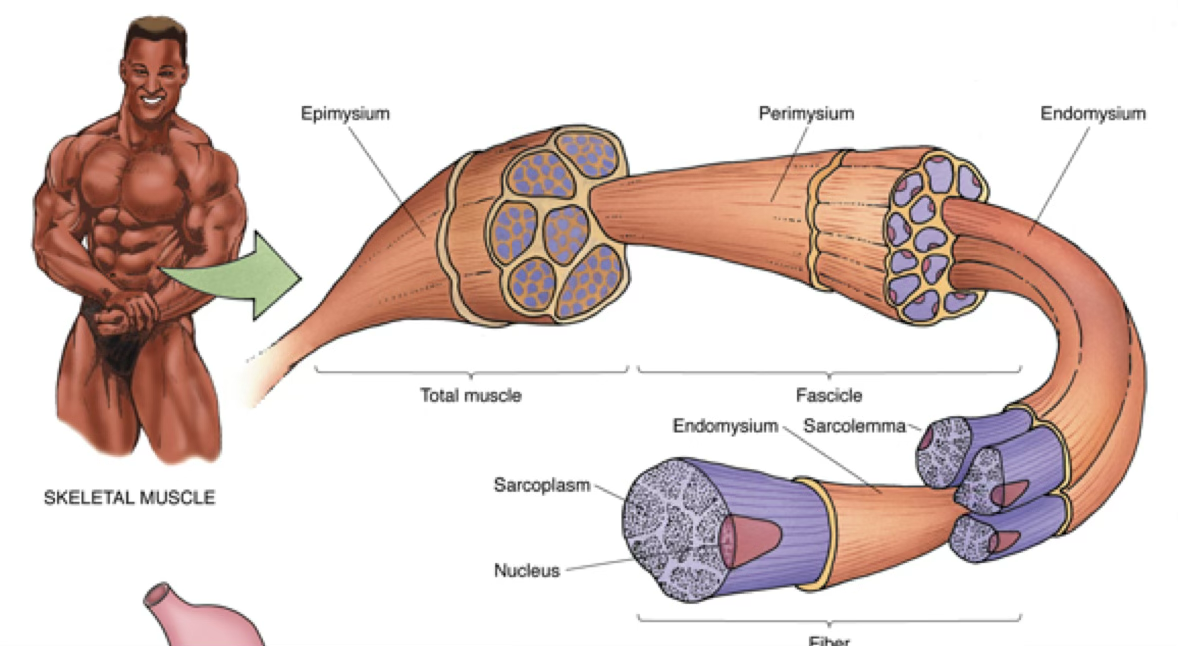
| Layer | Description | Surrounds |
|---|---|---|
| Endomysium | Basal lamina + reticular fibers | Individual muscle fibers |
| Perimysium | Less dense irregular CT | Bundles (fascicles) of fibers |
| Epimysium | Dense irregular CT. Outmost Layer. | Entire gross muscle |
Important
ALL muscle cells have basal laminae.
Anki cloze
The connective tissue layer surrounding individual muscle fibers is the {1:endomysium}; fascicles are surrounded by the {2:perimysium}; the entire muscle is covered by the {3:epimysium}.
Myofibrils and Sarcomeres
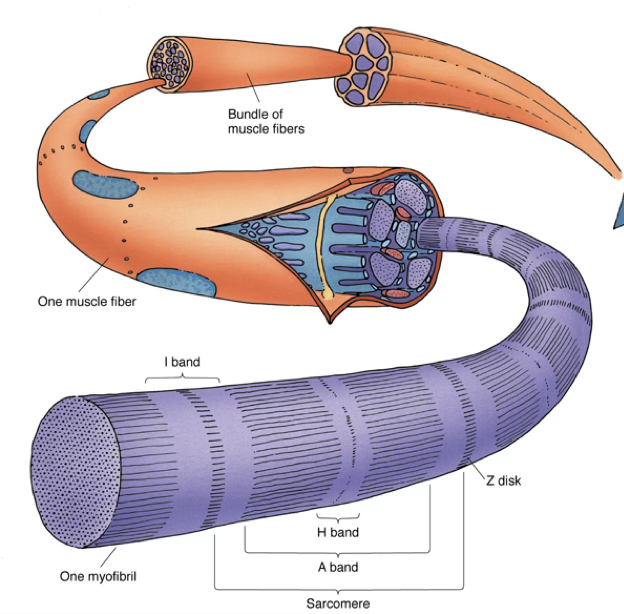
- Much of each muscle fiber is composed of longitudinal arrays of cylindrical myofibrils (1–2 µm in diameter)
- Myofibrils extend the entire length of the cell and are aligned precisely → responsible for cross-striations
The Sarcomere (contractile unit)
- Region between two successive Z disks
- Resting length: ~2.5 µm
| Band/Line | Description | What changes during contraction? |
|---|---|---|
| A band (dark) | Anisotropic; birefringent in polarized light; full length of myosin thick filaments | Width unchanged |
| I band (light) | Isotropic; actin only (no myosin overlap) | Narrows |
| H band | Center of A band; myosin tails only (no actin overlap) | Disappears |
| M line (Mittelscheibe) | Bisects H band | Stays |
| Z disk (Zwischenscheiben) | Bisects I band; anchors thin filaments | Moves closer together |
Mnemonic — Band names
“I Am Healthy, Mighty Z”
I band (light) → A band (dark) → H zone (lighter center) → M line → Z disk
Anki cloze
During muscle contraction, the {1:I band} and {2:H band} decrease in width, while the {3:A band} width remains unchanged.
Anki cloze
The sarcomere is defined as the region between two successive {1:Z disks} and has a resting length of approximately {2:2.5 µm}.
Classifying...
Myofilaments
Thick Filaments (myosin)
- Diameter: 15 nm; Length: 1.5 µm
- Composed of myosin II
- Form parallel arrays interdigitating with thin filaments
Thin Filaments (actin)
- Diameter: 7 nm; Length: 1.0 µm
- Composed primarily of F-actin (300–400 G-actin monomers), troponin, and tropomyosin
- Originate at the Z disk and project toward the center of two adjacent sarcomeres (opposite directions)
Anki cloze
Thick myofilaments are composed of {1:myosin} (diameter 15 nm), while thin myofilaments are composed primarily of {2:F-actin} (diameter 7 nm).
Sarcomere Structural Proteins
| Protein | Associated filament | Key function |
|---|---|---|
| Titin | Thick (myosin) | Controls thick filament assembly; sarcomere elasticity (spring between thick filament end and Z disk) |
| Nebulin | Thin (F-actin) | Stabilizes F-actin; maintains length of thin filaments; inserts into Z disk |
| Desmin | Z disk (intermediate filament, 10 nm) | Stabilizes myofibrils; links myofibrils to sarcolemma via costameres; positions mitochondria |
| Vimentin | Z disk | Co-localizes with desmin at Z disk |
| α-actinin | Z disk | Cross-links actin at Z disk |
| Tropomyosin | Thin filament groove | Blocks myosin binding site at rest |
| Troponin complex | Thin filament | Regulates actin–myosin interaction via Ca²⁺ |
Desmin functions in detail
- Stabilizes myofibrils — Desmin filaments encircle Z disks and link adjacent myofibrils via plectin.
- Links myofibrils to sarcolemma — Inserts into costameres; together with the dystrophin-associated protein complex, transduces contractile force from Z disk to basal lamina.
- Positions mitochondria — Loss of desmin → impaired mitochondrial positioning → cell death via energy deprivation or cytochrome c release.
Anki cloze
{1:Titin} is the giant protein that provides sarcomere elasticity by connecting the end of the thick filament to the Z disk.
Anki cloze
{1:Nebulin} is associated with thin filaments and acts as a stabilizer required for maintaining F-actin length.
Troponin–Tropomyosin Complex and Ca²⁺ Regulation
- Tropomyosin: two α-helical rods twisted together; sits in groove of F-actin near myosin head
- Troponin (Tn) = complex of three proteins:
- TnI — binds actin
- TnC — binds Ca²⁺ (found only in striated muscle)
- TnT — binds tropomyosin
Resting state: Ca²⁺ bound only to high-affinity site of TnC → tropomyosin blocks myosin binding site on F-actin.
Contraction: ↑ cytosolic Ca²⁺ → Ca²⁺ occupies low-affinity site of TnC → conformational change propagates through TnI → TnT → tropomyosin → myosin binding site exposed.
Anki cloze
In the troponin complex, {1:TnC} binds Ca²⁺, {2:TnI} binds actin, and {3:TnT} binds tropomyosin.
Anki cloze
TnC is found only in {1:striated muscle} (not smooth muscle, which uses calmodulin instead).
T Tubules and the Sarcoplasmic Reticulum
- T tubules: tubular invaginations of the sarcolemma that course transversely across the fiber
- Each sarcomere has two sets of T tubules, one at each A–I junction
- Conduct waves of depolarization deep into the cell interior
- Sarcoplasmic reticulum (SR): stores intracellular calcium; forms a meshwork around each myofibril
- Dilated terminal cisternae at each A–I junction
The Triad
One T tubule flanked by two terminal cisternae = triad (located at A–I junction in skeletal muscle)
Important
The triad arrangement allows a wave of depolarization to spread almost instantaneously from the sarcolemma to the terminal cisternae, which contain voltage-gated calcium release channels (junctional feet / ryanodine receptors).
Anki cloze
In skeletal muscle, a {1:triad} consists of one T tubule flanked by two {2:terminal cisternae} of the sarcoplasmic reticulum, located at the {3:A–I junction}.
Sliding Filament Theory (Huxley)
Core principle
Individual thick and thin filaments do not shorten during contraction. Instead, thin filaments slide past thick filaments, bringing Z disks closer together.
- Greater overlap between thick and thin filaments → I band and H band shrink
- A band width is constant (length of myosin thick filaments unchanged)
- Muscle shortens ~one-third of its original length
Anki cloze
According to Huxley’s sliding filament theory, during contraction the {1:thin} filaments slide past the {2:thick} filaments; filament lengths themselves do {3:not} change.
Skeletal Muscle Contraction — Sequence of Events
Step-by-step contraction cycle
First — Excitation:
- Motor neuron releases ACh (Ca²⁺-dependent) → electrical impulse generated in muscle
- Action potential travels across sarcolemma → down T tubules
- Depolarization activates dihydropyridine receptors (DHPR) on T tubule membrane
- DHPR activates ryanodine receptors on SR membrane → Ca²⁺ released from terminal cisternae into sarcoplasm
(excitation–contraction coupling occurs in ~15 ms)
Second — Contraction cycle:
- Ca²⁺ binds TnC → conformational change → tropomyosin shifts → myosin binding sites on actin exposed
- Energized myosin head (with ADP + Pᵢ bound) binds actin → crossbridge forms
- Pᵢ released → stronger actin–myosin bond; ADP released → power stroke (thin filament pulled toward center of sarcomere)
- New ATP binds S1 subfragment of myosin → crossbridge detaches
- ATP hydrolyzed → myosin head re-cocked (energized)
- Cycle continues while ATP is available and [Ca²⁺] in sarcoplasm is high
Rigor Mortis
- Begins 3–4 hours after death; lasts ~24–36 hours
- After death, Ca²⁺ leaks from SR → myosin heads bind actin
- No ATP synthesis → myosin cannot detach from actin
- Resolved only when proteolytic enzymes digest decomposing cells
Anki cloze
In the contraction cycle, ATP binding to myosin causes {1:detachment} of the crossbridge from actin, while ATP hydrolysis {2:re-cocks (energizes)} the myosin head.
Anki cloze
Rigor mortis occurs because after death, Ca²⁺ leaks from the SR allowing crossbridge formation, but absence of {1:ATP} prevents myosin detachment from actin.
Neuromuscular Junction (Motor End Plate)
Structure:
- Motor nerve fibers: myelinated; axon arborizes → loses myelin sheath
- Axon terminal contains: mitochondria, smooth ER, ~300,000 synaptic vesicles (40–50 nm diameter) containing acetylcholine (ACh)
- Axon terminal covered by Schwann cells
Transmission sequence:
- Action potential → opens voltage-gated Ca²⁺ channels in axon terminal
- Ca²⁺ → synaptic vesicle fusion → ACh released into synaptic cleft
- ACh binds nicotinic ACh receptors (ligand-gated ion channels) on sarcolemma → ion influx → depolarization + action potential
- Impulse spreads via T tubules → initiates contraction
- Acetylcholinesterase degrades ACh → acetate + choline → resting potential restored
- Choline recycled via Na⁺–choline symport into axon terminal → resynthesized into ACh by choline acetyltransferase
Anki cloze
ACh is degraded at the neuromuscular junction by {1:acetylcholinesterase} into acetate and choline, allowing the resting potential to be re-established.
Anki cloze
Each axon terminal contains approximately {1:300,000} synaptic vesicles, each 40–50 nm in diameter, filled with {2:acetylcholine}.
Muscle Regeneration — Skeletal Muscle
- Hypertrophy: increase in cell size
- Hyperplasia/regeneration: increase in cell number via satellite cells (resident stem cells)
Cardiac Muscle
General Characteristics
- Found only in the heart and pulmonary veins where they join the heart
- Elongated, branched cells
- 1 or 2 centrally placed nuclei (often with a biconical clear perinuclear area containing organelles, glycogen, and atrial natriuretic factor in atrial cells)
- Contraction: involuntary, vigorous, rhythmic
- Size: 10–15 µm diameter; 80–100 µm length
- Self-excitatory and electrically coupled (inherent rhythmicity)
- Rate modulated by autonomic NS — neuroendocrine innervation (no motor end plates)
Anki cloze
Cardiac muscle cells have {1:1 or 2} centrally placed nuclei and are joined end-to-end by {2:intercalated disks}.
Intercalated Disks
The key distinguishing feature of cardiac muscle.
| Component | Location in disk | Function |
|---|---|---|
| Fascia adherens | Transverse portion | Mechanical coupling; thin filaments anchor here (analogous to Z disk) |
| Macula adherens (desmosome) | Transverse portion | Mechanical coupling; resists traction forces |
| Gap junctions | Lateral portion | Electrical coupling; rapid propagation of action potential between cells |
Important
Gap junctions allow the myocardium to function as a functional syncytium — an action potential generated in one cell spreads rapidly to all coupled cells.
Anki cloze
The {1:transverse} portion of the intercalated disk contains fascia adherens and desmosomes (mechanical coupling), while the {2:lateral} portion contains gap junctions (electrical coupling).
Anki cloze
In cardiac muscle, the fascia adherens is analogous to the {1:Z disk} of skeletal muscle, as it anchors thin filaments.
Cardiac Muscle Ultrastructure
- T tubules: ~2× larger in diameter than in skeletal muscle; carry Ca²⁺ from extracellular space into cell at depolarization
- SR and T tubules form diads (not triads) located at Z lines (not A–I junction)
- SR does not form terminal cisternae → cannot store enough Ca²⁺ alone
- Negatively charged external lamina of T tubules stores Ca²⁺ for instantaneous release
- Nebulin extends up to only ~20% of the length of thin filaments (vs. full length in skeletal muscle)
- Numerous mitochondria: up to 40% of cell volume
Calcium dependence of cardiac muscle
Because SR Ca²⁺ stores are insufficient, cardiac muscle depends on extracellular Ca²⁺ entering via T tubules. Slow Na⁺/Ca²⁺ channels remain open for several tenths of a second, allowing extracellular Na⁺ and Ca²⁺ to enter and supplement internal stores.
Lack of extracellular Ca²⁺ → cessation of cardiac contraction within 1 minute (vs. skeletal muscle, which can continue for several hours).
Anki cloze
In cardiac muscle, T tubules and SR form {1:diads} located at the {2:Z line}, compared to triads at the A–I junction in skeletal muscle.
Anki cloze
Cardiac muscle stops contracting within {1:1 minute} if extracellular Ca²⁺ is removed, whereas skeletal muscle can continue for {2:several hours}.
Cardiac Connective Tissue
- Endomysium: surrounds each cardiac muscle cell (highly vascularized)
- Perimysium: surrounds fascicles
- Epicardium (≠ epimysium): connective tissue ensheathing the heart muscle
Cardiac Muscle Regeneration
- Previously thought to be entirely non-proliferative
- Post-infarction remodeling primarily by fibroblasts (fibrosis/scarring)
- New evidence: some mitotic cardiomyocytes and limited regeneration do occur
Smooth Muscle
General Characteristics
- Non-striated (contractile machinery present but irregularly arranged)
- Fusiform cells with a single, central, oval nucleus (2+ nucleoli)
- On contraction → nucleus assumes a corkscrew appearance
- Contraction: involuntary, slow, sustained
- Size: 0.2–2 µm diameter; 20–200 µm length
- No T tubule system
- Some smooth muscle capable of exogenous protein synthesis (collagen, elastin, glycosaminoglycans, proteoglycans, growth factors)
Locations of smooth muscle
- Walls of arteries and blood vessels
- Walls of hollow visceral organs (GI, urogenital)
- Walls of airways
- Dermis (arrector pili muscles)
- Ciliary muscle of the eye (lens focus)
Types of Smooth Muscle
| Type | Coupling | Nerve supply |
|---|---|---|
| Multiunit | Independent; each cell has its own nerve | Dense, individual innervation |
| Unitary (single-unit/vascular) | Electrically coupled via gap junctions; act in concert | Sparse; only a few cells directly innervated |
Smooth Muscle Ultrastructure
- Actin and myosin filaments (non-striated arrangement)
- Intermediate filaments: desmin (and vimentin in vascular smooth muscle)
- Dense bodies: membrane-associated and cytoplasmic; contain α-actinin (analogous to Z lines; actin filaments insert here)
- Caveolae: membrane invaginations forming a vesicular system continuous with SER — functionally analogous to sarcoplasmic reticulum; regulate cytosolic free Ca²⁺
- Microtubules present
Anki cloze
In smooth muscle, {1:dense bodies} are analogous to Z lines of striated muscle and serve as insertion points for actin filaments.
Anki cloze
Smooth muscle uses {1:caveolae} as a functionally analogous structure to the T tubules of striated muscle.
Smooth Muscle Contraction — Ca²⁺/Calmodulin Pathway
Key difference from striated muscle
- Smooth muscle thin filaments have no troponin
- Regulation is via Ca²⁺–calmodulin and phosphorylation of myosin light chain
- Myosin in resting state: actin-binding site masked by its light meromyosin moiety
Step-by-step:
- Stimulation (neurotransmitters, hormones, or autorhythmic signals) → Ca²⁺ released from caveolae/SER and/or enters via voltage-gated or ligand-gated channels
- Ca²⁺ binds calmodulin → conformational change of calmodulin
- Ca²⁺–calmodulin complex activates myosin light chain kinase (MLCK)
- MLCK phosphorylates the regulatory myosin light chain
- Phosphorylated light chain unmasks myosin’s actin-binding site → myosin unfolds into “golf club” shape → actin–myosin interaction → contraction
- Contraction continues as long as myosin is phosphorylated
- “Latch” state: myosin head attached to actin becomes dephosphorylated → decreased ATPase activity → myosin head cannot detach → sustained contraction with minimal ATP use
Mnemonic — Smooth Muscle Contraction
“Ca binds CAM, CAM calls MLCK, MLCK Phosphorylates, Phospho-chain Pulls”
Ca²⁺ → Calmodulin → MLCK → Phosphorylation → contraction
Anki cloze
In smooth muscle, Ca²⁺ binds {1:calmodulin}, which activates {2:myosin light chain kinase (MLCK)}, which phosphorylates the {3:regulatory myosin light chain}, unmasking the actin-binding site.
Anki cloze
The smooth muscle “latch” state occurs when the myosin head attached to actin becomes {1:dephosphorylated}, causing decreased ATPase activity and {2:sustained} contraction with minimal ATP use.
Anki cloze
Unlike striated muscle (which uses {1:troponin}), smooth muscle regulates actin–myosin interaction through phosphorylation of the {2:myosin light chain} via calmodulin and MLCK.
Smooth Muscle Regeneration
- Hypertrophy (increase in size)
- Hyperplasia (increase in number): smooth muscle cells are inherently proliferative
- e.g., uterine myometrium, vascular smooth muscle
- Vascular pericytes can also provide a source of smooth muscle cells
Comparative Summary Table
| Feature | Skeletal Muscle | Cardiac Muscle | Smooth Muscle |
|---|---|---|---|
| Striation | Yes | Yes | No |
| Nuclei | Multiple, peripheral | 1–2, central | 1, central |
| Cell shape | Long cylinder | Branched, elongated | Fusiform |
| Cell diameter | 10–100 µm | 10–15 µm | 0.2–2 µm |
| Cell length | Up to 30 cm | 80–100 µm | 20–200 µm |
| Control | Voluntary | Involuntary | Involuntary |
| Contraction type | Strong, quick, discontinuous | Strong, quick, continuous, rhythmic | Weak, slow, sustained |
| T tubules | Yes (at A–I junction) | Yes (at Z line, 2× wider) | No (caveolae instead) |
| SR | Well-developed with terminal cisternae | Sparse, no terminal cisternae | Sparse (caveolae/SER) |
| Ca²⁺ regulation | Troponin C | Troponin C + extracellular Ca²⁺ | Calmodulin + MLCK |
| Intercalated disks | No | Yes | No |
| Innervation | NMJ (motor end plate) | Neuroendocrine (autonomic) | Neuroendocrine (autonomic) |
| Regeneration | Via satellite cells | Limited (fibrosis predominant) | Proliferative (SMCs + pericytes) |
| Key structural unit | Sarcomere (Z–Z) | Sarcomere (Z–Z) | Dense bodies |
TLDR
Muscle Tissue — Full Summary
Embryology: All muscle from mesoderm. Three types: skeletal (voluntary, multinucleate), cardiac (involuntary, branched, central nuclei), smooth (involuntary, fusiform, non-striated).
Terminology: Sarcolemma = membrane; sarcoplasm = cytoplasm; sarcoplasmic reticulum = SER; sarcosomes = mitochondria.
Skeletal muscle organization (outer → inner): epimysium → perimysium (fascicles) → endomysium (individual fibers) → sarcolemma → myofibrils → sarcomeres.
Sarcomere structure (Z–Z): I band (actin only) | A band (myosin + actin overlap; constant width) | H zone (myosin tails only; disappears in contraction) | M line (center of H). On contraction: I band ↓, H band disappears, Z disks approximate; A band unchanged.
Filaments: Thick = myosin (15 nm, 1.5 µm). Thin = F-actin + troponin + tropomyosin (7 nm, 1.0 µm). Structural proteins: titin (sarcomere elasticity), nebulin (thin filament stabilizer), desmin (Z-disk integrity, costamere linkage, mitochondria positioning).
Ca²⁺ in striated muscle: SR releases Ca²⁺ → TnC binds Ca²⁺ → tropomyosin shifts → myosin binding site exposed → crossbridge cycle (bind actin, power stroke on Pᵢ release, detach on ATP binding, re-cock on ATP hydrolysis).
T-tubule system (skeletal): T tubule invaginations at A–I junctions; flanked by two terminal cisternae = triad; DHPR (T-tubule) activates ryanodine receptor (SR) → Ca²⁺ release.
NMJ: ACh released (Ca²⁺-dependent) → binds nicotinic receptors → depolarization → action potential → T tubules → Ca²⁺ release. ACh degraded by acetylcholinesterase; choline recycled by Na⁺–choline symport.
Rigor mortis: 3–4 h post-death; Ca²⁺ leak + no ATP → permanent crossbridges until proteolysis.
Cardiac muscle differences: Branched cells, intercalated disks (fascia adherens + desmosomes [transverse/mechanical] + gap junctions [lateral/electrical]). Diads at Z line (not triads at A–I). SR sparse → relies on extracellular Ca²⁺ via large T tubules + negatively charged external lamina. Loses contraction in ~1 min without extracellular Ca²⁺. Inherent rhythmicity; no motor end plates.
Smooth muscle: No troponin; no T tubules (caveolae instead); dense bodies = Z-line analogs. Regulation: Ca²⁺ → calmodulin → MLCK → myosin light chain phosphorylation → unmasking of actin-binding site → contraction. “Latch” state = dephosphorylated crossbridge → sustained low-ATP contraction. Two types: multiunit (independent) vs. unitary (gap junctions, act together).
Regeneration: Skeletal = satellite cells. Cardiac = mostly fibrosis (limited cardiomyocyte renewal). Smooth = proliferative SMCs + pericytes.